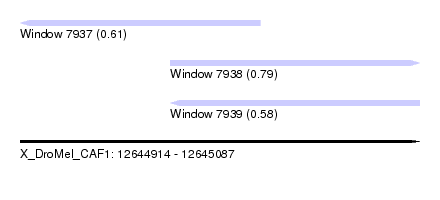

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,644,914 – 12,645,087 |
| Length | 173 |
| Max. P | 0.790619 |

| Location | 12,644,914 – 12,645,018 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.47 |
| Mean single sequence MFE | -25.87 |
| Consensus MFE | -17.95 |
| Energy contribution | -18.07 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.613014 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12644914 104 - 22224390 GAACAUAUUU-UUUGUAACCUACAUUUUUGGCAAAUGAAAGUUGCCGGGUGUUGUUGCAUCUUUUGCCAGCGAGCUCCUUUCAUUUCACUUUCCCAGAAAUAGU---------------A ..........-.........(((((((((((.(((((((((..((....(((((..(((.....)))))))).))..))))))))).......)))))))).))---------------) ( -22.10) >DroSim_CAF1 689 118 - 1 GAACAUAUUUUUUUGUAACCUACAUUUUUGGCAAAUGAAAGUUGCCGGGUGUUGUUGCAUCUCUUG-CAGCGAGCUCCUUUCAUUUUACUUACCCAGAAAUACAUACCCUAUUAC-CAGU ..(((........))).......((((((((.(((((((((.....((((.((((((((.....))-))))))))))))))))))).......))))))))..............-.... ( -24.10) >DroYak_CAF1 866 118 - 1 GAACAUAUUU-UUCGUAACCUACAUUUUUGGCAAAUGAAAGUUGCCGGGUGUUGUUGCAUCUUUUG-CAGCGCGCUCCUUUCAUUUUACUUGCCCCAAAAUGCAUACCCUAUUAUGCAUU ..........-..................((((((((((((.....((((((.((((((.....))-))))))))))))))))).....)))))....((((((((......)))))))) ( -31.40) >consensus GAACAUAUUU_UUUGUAACCUACAUUUUUGGCAAAUGAAAGUUGCCGGGUGUUGUUGCAUCUUUUG_CAGCGAGCUCCUUUCAUUUUACUUACCCAGAAAUACAUACCCUAUUA__CA_U .......................((((((((.(((((((((..(((((((((....))))).........)).))..))))))))).......))))))))................... (-17.95 = -18.07 + 0.11)
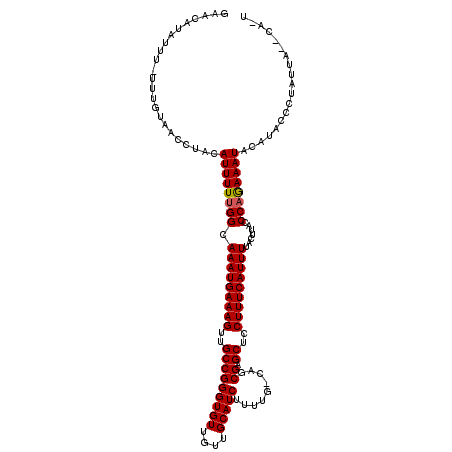


| Location | 12,644,979 – 12,645,087 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.70 |
| Mean single sequence MFE | -33.27 |
| Consensus MFE | -25.15 |
| Energy contribution | -24.93 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.790619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12644979 108 + 22224390 UUUCAUUUGCCAAAAAUGUAGGUUACAAA-AAAUAUGUUCCGCUGGCACUUGAGC---AGCCAAGUGGGUCGCUGUUCUAAGUGCUCAGUUGG--------GUUGUUUCAUUUUUUUUUU ...(((((.....)))))........(((-(((.(((.(((((((((((((((((---(((..........))))))).)))))).))).)))--------)......))).)))))).. ( -28.70) >DroSim_CAF1 767 106 + 1 UUUCAUUUGCCAAAAAUGUAGGUUACAAAAAAAUAUGUUCCGCUGGCACUUGAGC---AGCCAAGUGGGUCGCUGUUCUAAGUGCUCAGUUGG--------GUUUUU---UUUUGUUUUU ...(((((.....)))))......(((((((((...(..(.((((((((((((((---(((..........))))))).)))))).)))).).--------.).)))---)))))).... ( -33.00) >DroYak_CAF1 945 116 + 1 UUUCAUUUGCCAAAAAUGUAGGUUACGAA-AAAUAUGUUCCGCUGGCACUUGAGCAGCAGCCAAGUGGGUCGCUGUUCUAAGUGCUCAGUUGGGAAGGAGGGUUCUU---UCUUUUUUUU ((((....(((.........)))...)))-)........(((((((((((((((((((.(((.....))).))))))).)))))).))).)))(((((((((.....---))))))))). ( -38.10) >consensus UUUCAUUUGCCAAAAAUGUAGGUUACAAA_AAAUAUGUUCCGCUGGCACUUGAGC___AGCCAAGUGGGUCGCUGUUCUAAGUGCUCAGUUGG________GUUCUU___UUUUUUUUUU .((((..(((((.....((.((..(((........))).)))))))))..))))......((((.((((.((((......)))))))).))))........................... (-25.15 = -24.93 + -0.22)



| Location | 12,644,979 – 12,645,087 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.70 |
| Mean single sequence MFE | -25.73 |
| Consensus MFE | -16.77 |
| Energy contribution | -17.10 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.583251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12644979 108 - 22224390 AAAAAAAAAAUGAAACAAC--------CCAACUGAGCACUUAGAACAGCGACCCACUUGGCU---GCUCAAGUGCCAGCGGAACAUAUUU-UUUGUAACCUACAUUUUUGGCAAAUGAAA .....(((((((.......--------((..(((.((((((.((.((((.(......).)))---).))))))))))).)).(((.....-..)))......)))))))........... ( -24.10) >DroSim_CAF1 767 106 - 1 AAAAACAAAA---AAAAAC--------CCAACUGAGCACUUAGAACAGCGACCCACUUGGCU---GCUCAAGUGCCAGCGGAACAUAUUUUUUUGUAACCUACAUUUUUGGCAAAUGAAA ....((((((---(((...--------.(..(((.((((((.((.((((.(......).)))---).)))))))))))..)......)))))))))......(((((.....)))))... ( -25.20) >DroYak_CAF1 945 116 - 1 AAAAAAAAGA---AAGAACCCUCCUUCCCAACUGAGCACUUAGAACAGCGACCCACUUGGCUGCUGCUCAAGUGCCAGCGGAACAUAUUU-UUCGUAACCUACAUUUUUGGCAAAUGAAA ........((---((((.......((((...(((.((((((.((.(((((.((.....)).))))).))))))))))).))))....)))-)))........(((((.....)))))... ( -27.90) >consensus AAAAAAAAAA___AAAAAC________CCAACUGAGCACUUAGAACAGCGACCCACUUGGCU___GCUCAAGUGCCAGCGGAACAUAUUU_UUUGUAACCUACAUUUUUGGCAAAUGAAA ...........................(((((((.((((((((...(((.(......).)))....)).))))))))).((.(((........)))..)).......))))......... (-16.77 = -17.10 + 0.33)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:34:45 2006