| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,608,822 – 12,608,926 |
| Length | 104 |
| Max. P | 0.975242 |

| Location | 12,608,822 – 12,608,926 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 84.09 |
| Mean single sequence MFE | -25.32 |
| Consensus MFE | -18.40 |
| Energy contribution | -19.15 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.75 |
| SVM RNA-class probability | 0.975242 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12608822 104 - 22224390 AGGACCAGCAGGAAAAACAUGAAAUUGU---------AAUGGAGUCGAUAAAAAAAAAAUCAAUCUAUGUGAAUGGGUUUCGCAAUCCAGAGA-AAGAAGUUGGUCAAAUUUUU ..(((((((.......(((......)))---------..((((...(((.........)))......((((((.....)))))).))))....-.....)))))))........ ( -22.00) >DroSec_CAF1 6801 100 - 1 AGGACCAGCAGGAAAAACAUGUAAUUGU---------AAUGGAGUCGAUGA-----AAAUCAAUAUAUGUCUCUGGGUUUCGCAAUCCAGAGAGAAGAAGUUGGUCAAAUUUUU ..(((((((........((((((.....---------...(....)(((..-----..)))..))))))((((((((((....))))))))))......)))))))........ ( -29.40) >DroSim_CAF1 8222 100 - 1 AGGACCAGCAGGAAAAACAUGAAAUUGU---------AAUGGAGUCGAUGA-----AAAUCAAUAUAUGUGACUGGGUUUCGCAAUCCAGAGAGAAGAAGUUGGUCAAAUUUUU ..(((((((.......(((......)))---------..((((...(((..-----..)))......(((((.......))))).))))..........)))))))........ ( -23.10) >DroEre_CAF1 7199 105 - 1 UGGACCAGCAGGCAAAACAUGAACUCGAAUGAACAUGAAUGGAGUCGAUGG-----AAAUCAAUAUAUGUGAGUGGGUUUCACAAUCCAG-G---GGAAGUUGGUCAAAUUCUU ..(((((((........((((..(......)..))))..((((...(((..-----..)))......((((((.....)))))).)))).-.---....)))))))........ ( -26.80) >consensus AGGACCAGCAGGAAAAACAUGAAAUUGU_________AAUGGAGUCGAUGA_____AAAUCAAUAUAUGUGACUGGGUUUCGCAAUCCAGAGA_AAGAAGUUGGUCAAAUUUUU ..(((((((.(......).....................((((...(((.........)))......((((((.....)))))).))))..........)))))))........ (-18.40 = -19.15 + 0.75)

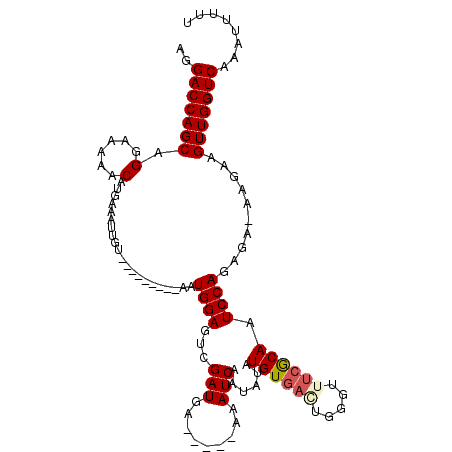

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:34:34 2006