
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,523,671 – 12,523,802 |
| Length | 131 |
| Max. P | 0.997765 |

| Location | 12,523,671 – 12,523,775 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.89 |
| Mean single sequence MFE | -36.68 |
| Consensus MFE | -14.19 |
| Energy contribution | -14.35 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.39 |
| SVM decision value | 2.93 |
| SVM RNA-class probability | 0.997765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12523671 104 - 22224390 CACCACGAUCAA---CGGUAUCGCGAUCUGACCCUGCAUCACCAUCAUCAU-ACACUGGUGC---AGCAGCAGAU------UGCGCAACAACAGCAUUAUCGUAGCCUCAAUGUG---GC ..(((((..(.(---(((((..((((((((...((((((((..........-....))))))---))...)))))------)))((.......))..)))))).)......))))---). ( -32.24) >DroSec_CAF1 53280 98 - 1 CACCACGAUCAG---CGGUAUCGCGAUCUGACCCUGCACCACC------AU-ACCCUGGUGC---AGCAGCAGAU------UGCGCAGCAACAGCAUUACCGUAGCCUCAAUGUG---GC ..(((((..(.(---(((((.(((((((((...((((((((..------..-....))))))---))...)))))------))))..((....))..)))))).)......))))---). ( -40.10) >DroEre_CAF1 53368 101 - 1 CACCACGACCAG---CGGUACCGCGACCUGACCCUGCACCACC---AUCAC-AGCCUGGUGC---AGCAGCAGAU------UGCGCAGCAACAGCACUACCGCAGCCUCAACGUG---GC ..(((((....(---(((((.(((((.(((...((((((((..---.....-....))))))---))...))).)------))))..((....))..))))))........))))---). ( -37.80) >DroYak_CAF1 53641 104 - 1 CACCACGAUCAG---CGGUAUCGCGAUCUAACGCUGCACCAUCAUCAUCAU-ACCCUCGUGC---AGCAGCAGAU------UGCGCAGCAGCAGCAUUACCGCAGCCUCAAUGUG---GC ...........(---(((((.((((((((...(((((((............-......))))---)))...))))------))))..((....))..)))))).(((.......)---)) ( -35.37) >DroAna_CAF1 63742 116 - 1 CACCACGAGCUGUCCCGGUACCGUGAUAUAGCCAUGCACCAGCAA-AUGGCCGCCAUGGUCCAUCAGCAGCAGCUGGUCCAGGCCCAGCACCAGCAGUACCGGCACC---AUCUGAGCGC ........(((((.(((((((.((......(((((((....))).-.)))).(((.(((...((((((....)))))))))))).........)).))))))).)).---.....))).. ( -37.90) >consensus CACCACGAUCAG___CGGUAUCGCGAUCUGACCCUGCACCACCA__AUCAU_ACCCUGGUGC___AGCAGCAGAU______UGCGCAGCAACAGCAUUACCGCAGCCUCAAUGUG___GC ...((((........(((((..((...........((((((...............))))))....((.((...........)))).......))..))))).........))))..... (-14.19 = -14.35 + 0.16)

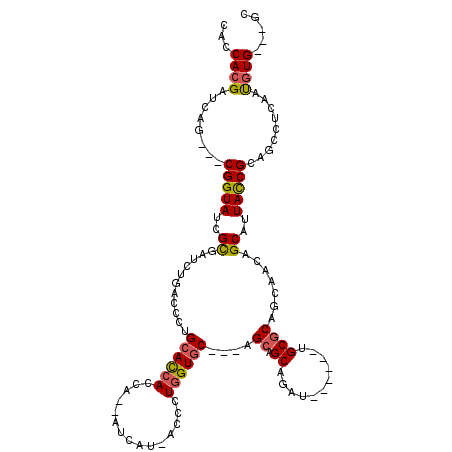

| Location | 12,523,707 – 12,523,802 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 77.51 |
| Mean single sequence MFE | -34.57 |
| Consensus MFE | -14.47 |
| Energy contribution | -14.47 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.42 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.793488 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12523707 95 - 22224390 GUGGCGCAGCCACCGCC---GCCGACCGCACACCACGAUCAA---CGGUAUCGCGAUCUGACCCUGCAUCACCAUCAUCAU-ACACUGGUGC---AGCAGCAGAU----- (((((...)))))...(---((..((((..............---))))...)))(((((...((((((((..........-....))))))---))...)))))----- ( -28.28) >DroSec_CAF1 53316 89 - 1 GUGGCCCAGCCACCGCC---GCCGACCGCACACCACGAUCAG---CGGUAUCGCGAUCUGACCCUGCACCACC------AU-ACCCUGGUGC---AGCAGCAGAU----- (((((...)))))...(---((..(((((............)---))))...)))(((((...((((((((..------..-....))))))---))...)))))----- ( -36.10) >DroEre_CAF1 53404 92 - 1 GUGGCGCAGCCGCCGCC---GCCGACCGCUCACCACGACCAG---CGGUACCGCGACCUGACCCUGCACCACC---AUCAC-AGCCUGGUGC---AGCAGCAGAU----- ((((((....))))))(---((.(((((((..........))---)))).).)))..(((...((((((((..---.....-....))))))---))...)))..----- ( -36.90) >DroYak_CAF1 53677 95 - 1 GUUGCGCAGCCACCGCC---GCCGACCGCACACCACGAUCAG---CGGUAUCGCGAUCUAACGCUGCACCAUCAUCAUCAU-ACCCUCGUGC---AGCAGCAGAU----- ((((.((.((....)).---)))))).((.......((((.(---((....)))))))....(((((((............-......))))---))).))....----- ( -28.07) >DroAna_CAF1 63779 109 - 1 GUGGGACAGCCACCGCCACCACCGGCCGCACACCACGAGCUGUCCCGGUACCGUGAUAUAGCCAUGCACCAGCAA-AUGGCCGCCAUGGUCCAUCAGCAGCAGCUGGUCC ..((((((((....(((......)))((.......)).))))))))((.((((((.....(((((((....))).-.))))...))))))))((((((....)))))).. ( -43.50) >consensus GUGGCGCAGCCACCGCC___GCCGACCGCACACCACGAUCAG___CGGUAUCGCGAUCUGACCCUGCACCACCA__AUCAU_ACCCUGGUGC___AGCAGCAGAU_____ (((((...))))).((....((..((((.................))))...))...........((((((...............))))))....))............ (-14.47 = -14.47 + -0.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:51 2006