
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,466,711 – 12,466,873 |
| Length | 162 |
| Max. P | 0.976606 |

| Location | 12,466,711 – 12,466,816 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 85.54 |
| Mean single sequence MFE | -34.33 |
| Consensus MFE | -27.42 |
| Energy contribution | -28.28 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.579325 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12466711 105 + 22224390 CCGUGUAUGUAUA-ACACGUGCCG---UGGAAACUGCGCGGAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUUC-CUA .((.((((((...-..))))))))---..(((((((.(((....)))...)))))))........(((....)))..(((((((((....))))))))).......-... ( -36.30) >DroSec_CAF1 5542 105 + 1 CCGUGUAUGUAUA-ACACGUGCCG---UGGAAACUGCGCGGAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAAUCGACGCGUUCGCACGCGUUGUUUUUUCC-CUA .((.((((((...-..))))))))---..(((((((.(((....)))...)))))))........(((....)))...((((((((....))))))))........-... ( -34.30) >DroSim_CAF1 5560 105 + 1 CCGUGUAUGUAUA-ACACGUGCCG---UGGAAACUGCGCGGAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAAUCGACGCGUUCGCACGCGUUGAUUUUUCC-CUA .((.((((((...-..))))))))---..(((((((.(((....)))...)))))))........(((....)))(((((((((((....))))))))))).....-... ( -37.90) >DroEre_CAF1 5582 106 + 1 CCGUGUAUGUAUA-ACACGUGCCG---UGGAAACUGCGCGGAAACGCUGCCAGUUUCAGUUUUUCGCUGUAGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUCCCCCA .((.((((((...-..))))))))---..(((((((((((....))).).)))))))........(((....)))..(((((((((....)))))))))........... ( -37.20) >DroYak_CAF1 5582 106 + 1 CCGUGUAUGUAUA-ACACGUGUCG---UGGAAACUGCGCGGAAACGCUGCCAGUUUCAGUUUUUCGCUGUAGAGCAUUCAACGCGCUUGCACGCGUUGAUUUUUCCCCCA .(((((.......-))))).....---..(((((((((((....))).).)))))))........(((....)))..((((((((......))))))))........... ( -34.90) >DroAna_CAF1 8797 104 + 1 UAUUUCAUAUACAUACACGUGCAGGAAAGGAAACUGCGCGA---GUUUUCCAGU-UCAGUU-UUCGGUGCCGAGCGUUCGACGCGUUCACACGCGUUUCUCCCCCC-CAA ................((((...((...((((((((.((..---........))-.)))))-)))....))..))))..(((((((....))))))).........-... ( -25.40) >consensus CCGUGUAUGUAUA_ACACGUGCCG___UGGAAACUGCGCGGAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUCC_CUA .((.((((((......)))))))).....(((((((.(((....)))...)))))))........(((....)))..((((((((......))))))))........... (-27.42 = -28.28 + 0.87)

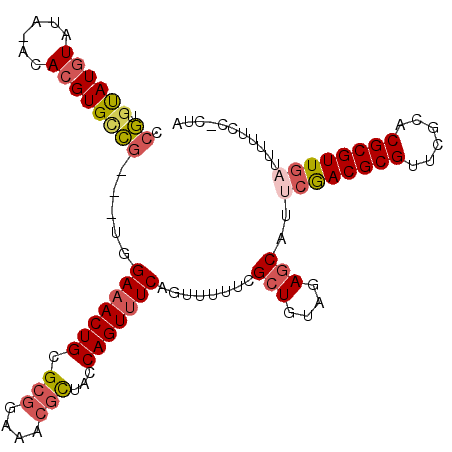

| Location | 12,466,747 – 12,466,846 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 92.81 |
| Mean single sequence MFE | -21.54 |
| Consensus MFE | -20.76 |
| Energy contribution | -20.48 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.915398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12466747 99 + 22224390 GAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUUC-CUAAAUCC-UAAAAAAGAACCCACAACCAUUAAG (((((.......))))).((((((.(((....)))..(((((((((....))))))))).......-........-.....)))))).............. ( -21.70) >DroSec_CAF1 5578 99 + 1 GAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAAUCGACGCGUUCGCACGCGUUGUUUUUUCC-CUAAAUCC-UAAAAAAGAACCCACAACCAAUAAG (((((.......))))).((((((.(((....)))...((((((((....))))))))........-........-.....)))))).............. ( -19.70) >DroSim_CAF1 5596 99 + 1 GAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAAUCGACGCGUUCGCACGCGUUGAUUUUUCC-CUAAAUCC-UAAAAAAGAACCCACAACCAAUAAG (((((.......))))).((((((.(((....)))(((((((((((....))))))))))).....-........-.....)))))).............. ( -23.30) >DroEre_CAF1 5618 101 + 1 GAAACGCUGCCAGUUUCAGUUUUUCGCUGUAGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUCCCCCAAAUUCUUAAAAGAGAACCCACAAACAUUAAG ((((.((((.......)))).))))(((....)))..(((((((((....))))))))).............(((((....)))))............... ( -22.50) >DroYak_CAF1 5618 100 + 1 GAAACGCUGCCAGUUUCAGUUUUUCGCUGUAGAGCAUUCAACGCGCUUGCACGCGUUGAUUUUUCCCCCAAAUCC-UAAAAAAGAAUCCACAAACAAUAAG (((((.......))))).((..((((((....)))..((((((((......))))))))................-.......)))...)).......... ( -20.50) >consensus GAAACGCUACCAGUUUCAGUUUUUCGCUGUAGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUCC_CUAAAUCC_UAAAAAAGAACCCACAACCAAUAAG (((((.......))))).((((((.(((....)))..((((((((......))))))))......................)))))).............. (-20.76 = -20.48 + -0.28)

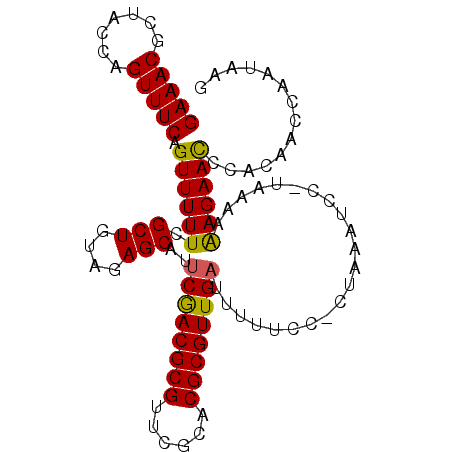

| Location | 12,466,777 – 12,466,873 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 92.40 |
| Mean single sequence MFE | -14.71 |
| Consensus MFE | -13.38 |
| Energy contribution | -13.75 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.77 |
| SVM RNA-class probability | 0.976606 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12466777 96 + 22224390 AGAGCAUUCGACGCGUUCGCACGCGUUGAUUUUUUC-CUAAAUCCUAAAAAAGAACCCACAACCAUUAAGCCAAAAUCUAUAAAAAAAAAUUGCAAA ...(((.(((((((((....)))))))))((((((.-.((.....))..))))))....................................)))... ( -14.50) >DroSec_CAF1 5608 96 + 1 AGAGCAAUCGACGCGUUCGCACGCGUUGUUUUUUCC-CUAAAUCCUAAAAAAGAACCCACAACCAAUAAGCCAAAAUCUAUAAAAAAAAAUUGCAAA ...(((((.(((((((....)))))))(((((((..-............))))))).................................)))))... ( -14.54) >DroSim_CAF1 5626 95 + 1 AGAGCAAUCGACGCGUUCGCACGCGUUGAUUUUUCC-CUAAAUCCUAAAAAAGAACCCACAACCAAUAAGCCAAAAUCUAUAAAA-AAAUUUGCAAA .(((.(((((((((((....))))))))))).))).-................................................-........... ( -16.50) >DroYak_CAF1 5648 97 + 1 AGAGCAUUCAACGCGCUUGCACGCGUUGAUUUUUCCCCCAAAUCCUAAAAAAGAAUCCACAAACAAUAAGCCAAAAUCUAUAUAAAAAAUUUGCAAA ...(((.((((((((......))))))))...............((.....))......................................)))... ( -13.30) >consensus AGAGCAAUCGACGCGUUCGCACGCGUUGAUUUUUCC_CUAAAUCCUAAAAAAGAACCCACAACCAAUAAGCCAAAAUCUAUAAAAAAAAAUUGCAAA .(((.((((((((((......)))))))))).))).............................................................. (-13.38 = -13.75 + 0.37)

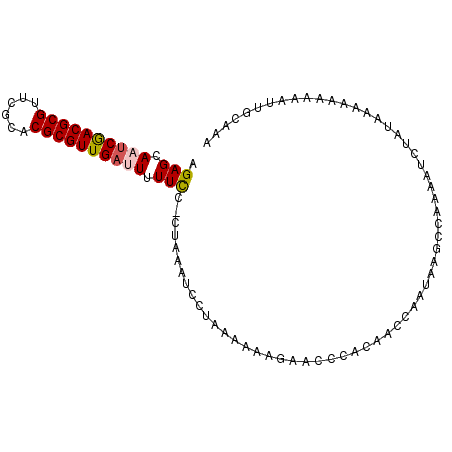
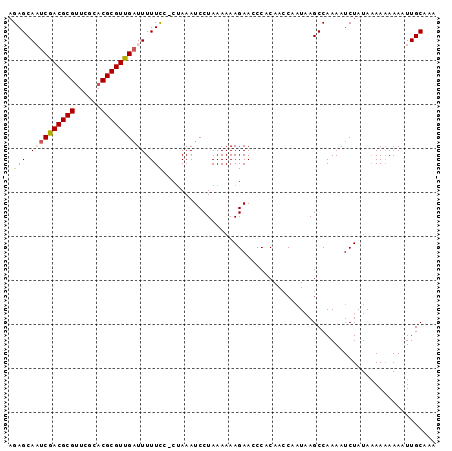
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:19 2006