
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,452,280 – 12,452,409 |
| Length | 129 |
| Max. P | 0.855449 |

| Location | 12,452,280 – 12,452,376 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 70.17 |
| Mean single sequence MFE | -23.27 |
| Consensus MFE | -12.24 |
| Energy contribution | -11.80 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.705185 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
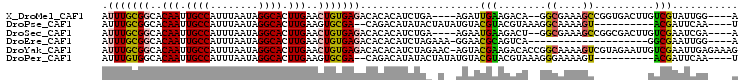
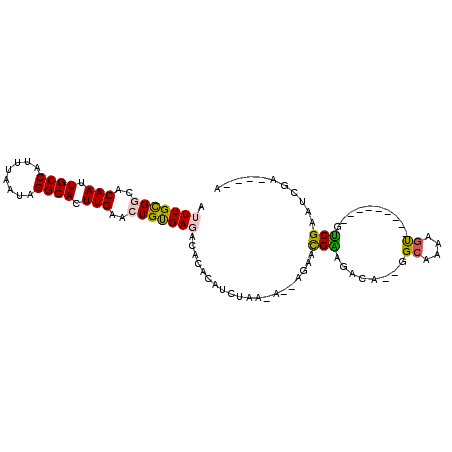
>X_DroMel_CAF1 12452280 96 - 22224390 AUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUGA----AGAUUGAAGACA--GGCGAAAGCCGGUGACUUGUCGUAUUGG----A .((..(((..(((.((((........)))).)))..)))..))((.((.(((...----.)))))..((((--(((....)))(....).)))))).....----. ( -28.00) >DroPse_CAF1 7087 90 - 1 AUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAAGUGCGA--CAGACAUAUACUAUAUGUACGUACGUAAAGGCAAAAGU----------ACGAUUCAA----U .(((((((((....))))............(((...(((((.--...((((((...)))))).)))))...))))))))...----------.........----. ( -21.50) >DroSec_CAF1 5345 96 - 1 AUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUGA----AGAAUGAAGACU--GGCGAAAGCCGGCGACUUGUCGAAUCGA----A ..(..(((..(((.((((........)))).)))..)))..)((((..((((...----.).)))..(.((--(((....))))))....)))).......----. ( -27.70) >DroEre_CAF1 5815 81 - 1 AUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUAGAAA-GGAACGCAGUCA--------------------GGCGAAUUGG----A ((((((....(((.((((........)))).))).((((((.(.....).(((.....-))).))))))..--------------------.))))))...----. ( -17.00) >DroYak_CAF1 5951 105 - 1 AUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUAGAAC-AGUACGAAGACACCGGCAAAAGUCGUAGAAUUGUCGAAUUGAGAAAG .((((((((.(((.((((........)))).))).(((((.(((......)))...))-)))..................)))))))).................. ( -23.70) >DroPer_CAF1 7095 90 - 1 AUUUGUGGCACAAUUGCCAUUUAAUAGGCACUUGAAGUGCGA--CAGACAUAUACUAUAUGUACGUACGUAAAGGGAAAAGU----------ACGAUUCAA----U ..((((.((((((.((((........)))).))...)))).)--)))((((((...)))))).(((((............))----------)))......----. ( -21.70) >consensus AUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUAA_A__AGAACGAAGACA__GGCAAAAGU_________GUCGAAUCGA____A .(((((((..(((.((((........)))).)))..)))))))....................(((........((....))..........)))........... (-12.24 = -11.80 + -0.44)

| Location | 12,452,312 – 12,452,409 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 84.18 |
| Mean single sequence MFE | -23.18 |
| Consensus MFE | -19.74 |
| Energy contribution | -19.60 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.855449 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
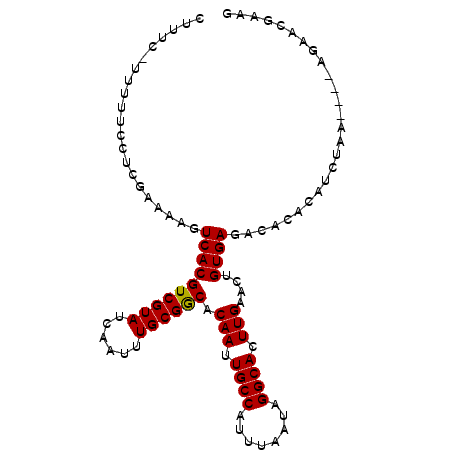
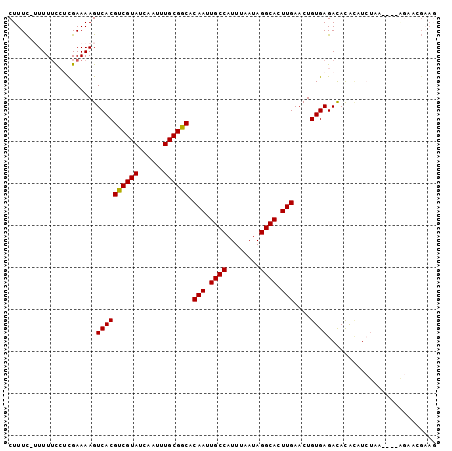
>X_DroMel_CAF1 12452312 97 - 22224390 CUUUC-AUUUUUCUCGAAAAGUCACGUCGUAUCAAUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUGA----AGAUUGAAG ..(((-(.(((((........((((((((((......)))))).(((.((((........)))).)))....))))((......)).))----))).)))). ( -24.50) >DroVir_CAF1 8201 86 - 1 CUUUUAUUUUUCGAUGAAAAGUCACGUCGUAUCAAUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGAUAUGAAUA---------------- .(((((((....)))))))..((((((((((......)))))).(((.((((........)))).)))....))))..........---------------- ( -20.90) >DroSec_CAF1 5377 97 - 1 CUUUC-AUUUUCCUCGAAAAGUCACGUCGUAUCAAUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUGA----AGAAUGAAG ..(((-(((((.((((...(((...((((((......)))))).(((.((((........)))).))).))).)))).((......)).----)))))))). ( -23.80) >DroEre_CAF1 5829 100 - 1 CUUUC-UUUUUCCUCGAAAAGUCACGUCGUAUCAAUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUAGAAA-GGAACGCAG (((((-((((((...))))).((((((((((......)))))).(((.((((........)))).)))....))))...........)))))-)........ ( -24.30) >DroWil_CAF1 29717 96 - 1 CUUUC-UUUUGC--UGAAAAGUCACGUCGUAUCAAUUUGCGACACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGAUAUAUAUAUAU---AUAUACGAAC .((((-......--.))))..((((((((((......)))))).(((.((((........)))).)))....)))).((((((....))---))))...... ( -22.20) >DroYak_CAF1 5989 100 - 1 CUUUC-UUUUUCCUCGAAAAGUCACGUCGUAUCAAUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUAGAAC-AGUACGAAG .....-.......(((.........((((((......)))))).(((.((((........)))).))).(((((.(((......)))...))-))).))).. ( -23.40) >consensus CUUUC_UUUUUCCUCGAAAAGUCACGUCGUAUCAAUUUGCGGCACAAUUGCCAUUUAAUAGGCACUUGAACUGUGAGACACACAUCUAA____AGAACGAAG .....................((((((((((......)))))).(((.((((........)))).)))....)))).......................... (-19.74 = -19.60 + -0.14)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:14 2006