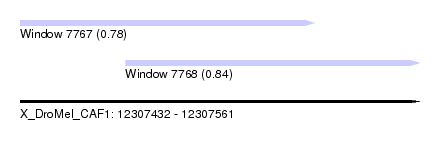
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,307,432 – 12,307,561 |
| Length | 129 |
| Max. P | 0.841069 |

| Location | 12,307,432 – 12,307,527 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 96.00 |
| Mean single sequence MFE | -25.26 |
| Consensus MFE | -24.84 |
| Energy contribution | -24.36 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.783320 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12307432 95 + 22224390 GCCAGAAAAUGUCCAAGCGUUCAGCACCAUUAGAUCGUUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGC ................((((.((((.(((((((....)))))))..............(((((((....)))..)))).........)))))))) ( -20.00) >DroSec_CAF1 9613 95 + 1 GCCAGAAAAUGGCCAAGCGCUCAGCACCACUAGAUCGCUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGC ((((.....))))...(((.(((((.(((((((....)))))))..............(((((((....)))..)))).........)))))))) ( -30.00) >DroSim_CAF1 10158 95 + 1 GCCAGAAAAUGGCCAAGCGCUCAGCACCACUAGAUCGCUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGC ((((.....))))...(((.(((((.(((((((....)))))))..............(((((((....)))..)))).........)))))))) ( -30.00) >DroEre_CAF1 9649 95 + 1 GCCAGAAAAUGGCCAAGCGCUCAGCACCCCUAGAUCGUUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGC ((((.....))))...(((.(((((.......((((((..(((.....))).......))))))......(((....))).......)))))))) ( -23.30) >DroYak_CAF1 9033 95 + 1 GCCAGAAAAUGGCCAAGUGCUCAGCAUCACUAGAUCGUUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGUUAUUCCCCAGCUGAUGC ((((.....)))).....(((((((.......((((((..(((.....))).......))))))......((((.....))))....))))).)) ( -23.00) >consensus GCCAGAAAAUGGCCAAGCGCUCAGCACCACUAGAUCGUUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGC ((((.....))))...(((.(((((.(((((((....)))))))..............(((((((....)))..)))).........)))))))) (-24.84 = -24.36 + -0.48)



| Location | 12,307,466 – 12,307,561 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 98.25 |
| Mean single sequence MFE | -21.58 |
| Consensus MFE | -21.76 |
| Energy contribution | -21.32 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.95 |
| Structure conservation index | 1.01 |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.841069 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12307466 95 + 22224390 UCGUUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGCUGCUUCUUCUGCUUCUGUUUCCCAUUUACAGCUC ..((..((((.(((((........(((((((....)))..)))).......((((....)))).............))))).))))..))..... ( -21.00) >DroSec_CAF1 9647 95 + 1 UCGCUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGCUGCUUCUUCUGCUUCUGUUUCCCAUUUACAGCUC ..((((((((.(((((........(((((((....)))..)))).......((((....)))).............))))).))))...)))).. ( -23.10) >DroSim_CAF1 10192 95 + 1 UCGCUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGCUGCUUCUUCUGCUUCUGUUUCCCAUUUACAGCUC ..((((((((.(((((........(((((((....)))..)))).......((((....)))).............))))).))))...)))).. ( -23.10) >DroYak_CAF1 9067 95 + 1 UCGUUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGUUAUUCCCCAGCUGAUGCUGCUUCUUCUGCUUCUGUUUCCCAUUUACAGCUC ..((..((((.(((((....(((((...(((....)))...))))).....((((....)))).............))))).))))..))..... ( -19.10) >consensus UCGCUGGUGGCAAACAUAAUUAAUGCGAUCAUCUUUGAGUUCGCUUUUCCCCAGCUGAUGCUGCUUCUUCUGCUUCUGUUUCCCAUUUACAGCUC ..((((((((.(((((........(((((((....)))..)))).......((((....)))).............))))).))))...)))).. (-21.76 = -21.32 + -0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:08 2006