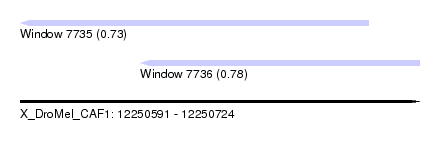
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,250,591 – 12,250,724 |
| Length | 133 |
| Max. P | 0.776963 |

| Location | 12,250,591 – 12,250,707 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 97.84 |
| Mean single sequence MFE | -41.20 |
| Consensus MFE | -37.01 |
| Energy contribution | -37.08 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.731564 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12250591 116 - 22224390 UUCUCUGUUGCAGCAGGGCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCACCGCCUGCGGUUGUUCUUACUCGGGAUUAAGACGCUUUUC ..((((((....))))))((((((.(((((((..((.(((........))).))..))))))).))))))(((((((((....)))).(((((.....)))))...)))))..... ( -40.10) >DroSec_CAF1 18616 115 - 1 UUCGCUGUUGCAGCAG-GCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCACCGCCUGCGGUUGUUCUUACUCGGGAUUAAGACGCUUUUC .........((.((((-((.((((((.(((((..(((((((((.....)))....))))))..)))))))))))....)))))).(((..(((......)))....)))))..... ( -41.30) >DroEre_CAF1 13854 115 - 1 UUCGCUGUAGCAGCCG-GCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCACCGCCUGCGGUUGUUCUUACUCGGGAUUAAGACGCUUUUC ...((....((((.((-(..((((((.(((((..(((((((((.....)))....))))))..)))))))))))..))).)))).(((..(((......)))....)))))..... ( -42.10) >DroYak_CAF1 7045 115 - 1 UUCGCUGUUGCAGCAG-GCCAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCACCGCCUGCGGUUGUUCUUACUCGGGAUUAAGACGCUUUUC .........((.((((-((.((((((.(((((..(((((((((.....)))....))))))..)))))))))))....)))))).(((..(((......)))....)))))..... ( -41.30) >consensus UUCGCUGUUGCAGCAG_GCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCACCGCCUGCGGUUGUUCUUACUCGGGAUUAAGACGCUUUUC ...((((...))))....((((((.(((((((..((.(((........))).))..))))))).))))))(((((((((....)))).(((((.....)))))...)))))..... (-37.01 = -37.08 + 0.06)



| Location | 12,250,631 – 12,250,724 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 96.20 |
| Mean single sequence MFE | -31.52 |
| Consensus MFE | -26.58 |
| Energy contribution | -26.82 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.776963 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12250631 93 - 22224390 GUGGAAUGAGCAAGUUUUUCUCUGUUGCAGCAGGGCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCA ...................((((((....))))))..((((((.(((((..(((((((((.....)))....))))))..))))))))))).. ( -30.40) >DroSec_CAF1 18656 91 - 1 GUGGGAUGAGCAAGUU-UUCGCUGUUGCAGCAG-GCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCA .((.(((.(((.....-...)))))).))...(-((.....((.(((((..(((((((((.....)))....))))))..))))).)).))). ( -30.90) >DroEre_CAF1 13894 91 - 1 GUGGGAUGAGCAAGUU-UUCGCUGUAGCAGCCG-GCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCA (((..(.........)-..)))((((((.....-)))))).((.(((((..(((((((((.....)))....))))))..))))).))..... ( -31.90) >DroYak_CAF1 7085 91 - 1 GUGGGAUGAGCAAGUU-UUCGCUGUUGCAGCAG-GCCAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCA (((.((((.((((.((-...((..((((((((.-(((((((.....))))))).))))))))...)))).)))).)))).))).......... ( -32.90) >consensus GUGGGAUGAGCAAGUU_UUCGCUGUUGCAGCAG_GCUAUAUCUGACAUGUGGCAUGCUGCAAGUUGCAAGUUGCAUGUUGCAUGUGGGUGUCA ....................((((...))))......((((((.(((((..(((((((((.....)))....))))))..))))))))))).. (-26.58 = -26.82 + 0.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:31:40 2006