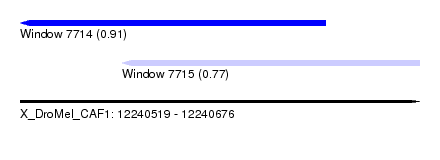
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,240,519 – 12,240,676 |
| Length | 157 |
| Max. P | 0.911321 |

| Location | 12,240,519 – 12,240,639 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.61 |
| Mean single sequence MFE | -33.24 |
| Consensus MFE | -22.68 |
| Energy contribution | -23.47 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911321 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12240519 120 - 22224390 AAAACCAUUUGAUAUUAAGAAGUCCAUUCGACCCUAUCAUUUGAUCAAAAGUGGUAGGGUCCUUAUGCUAACGGCCAAUCAAGUUAAUGCACUUACUCAAAAAGGCAAGGAAAAGGUGGG ...((((((((((.....(((.....)))(((((((((((((......)))))))))))))................)))))))...(((.(((.......)))))).......)))... ( -31.10) >DroSim_CAF1 63812 120 - 1 AAAACCCUUUGAUAUUAAGCAGUCCAUUCGACCCUAUCAUUUGAUCAAAAGUGGUAGGGUCAUUAUACUAACGGCCAAACAAGCUGAAGCACUCACUCAAAAAUGCAAUAAAAAGGUGGG ....((((((..((((..(((........(((((((((((((......)))))))))))))..........((((.......)))).................)))))))..)))).)). ( -32.10) >DroYak_CAF1 47878 120 - 1 AAGUCCCUACGAUAUUAAGAAGUCCACUCGACCCCAUGAUUUGAUCAAAAGUGGUAGGGUCAUUAUGCUAACCGCGAAUCAAGUGGAUGCAUUCACUCAAAGGGGCACGCAGAAUGUGCG ..((((((..((.........(((((((.(((((.((.((((......)))).)).)))))....(((.....))).....)))))))........))..)))))).((((.....)))) ( -36.53) >consensus AAAACCCUUUGAUAUUAAGAAGUCCAUUCGACCCUAUCAUUUGAUCAAAAGUGGUAGGGUCAUUAUGCUAACGGCCAAUCAAGUUGAUGCACUCACUCAAAAAGGCAAGAAAAAGGUGGG .......................((((..(((((((((((((......)))))))))))))....(((...((((.......))))..)))........................)))). (-22.68 = -23.47 + 0.78)

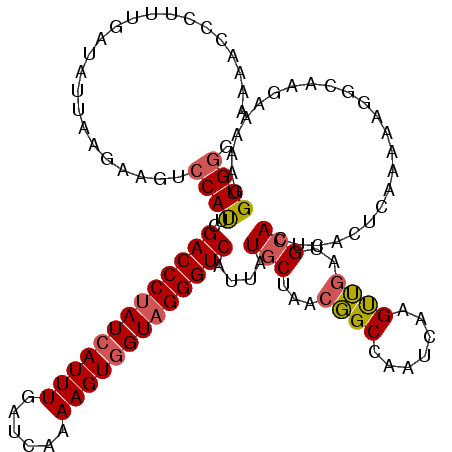
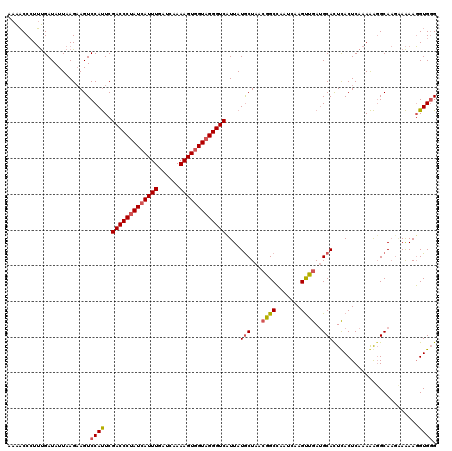
| Location | 12,240,559 – 12,240,676 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.47 |
| Mean single sequence MFE | -31.20 |
| Consensus MFE | -24.27 |
| Energy contribution | -25.27 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.772718 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12240559 117 - 22224390 GUGGCGAUGUCCCCA---AAAGUACGAUAUCCUAUACCGGAAAACCAUUUGAUAUUAAGAAGUCCAUUCGACCCUAUCAUUUGAUCAAAAGUGGUAGGGUCCUUAUGCUAACGGCCAAUC .((((((((((....---.......))))))......((......(((..(.......(((.....)))(((((((((((((......))))))))))))))..)))....))))))... ( -31.70) >DroSim_CAF1 63852 117 - 1 GUGGCGAUUAUCCCA---UAAGUACGAUAUCCUAUACUGGAAAACCCUUUGAUAUUAAGCAGUCCAUUCGACCCUAUCAUUUGAUCAAAAGUGGUAGGGUCAUUAUACUAACGGCCAAAC .((((..........---...((..((((((.......((.....))...))))))..)).........(((((((((((((......)))))))))))))............))))... ( -31.10) >DroYak_CAF1 47918 120 - 1 CUAGCAAUUCGCCUGGUGGUAGUACGAUAUCCUAUACCGGAAGUCCCUACGAUAUUAAGAAGUCCACUCGACCCCAUGAUUUGAUCAAAAGUGGUAGGGUCAUUAUGCUAACCGCGAAUC ......((((((..((((((((...(((.(((......))).))).))))(((........))))))).(((((.((.((((......)))).)).)))))............)))))). ( -30.80) >consensus GUGGCGAUUACCCCA___AAAGUACGAUAUCCUAUACCGGAAAACCCUUUGAUAUUAAGAAGUCCAUUCGACCCUAUCAUUUGAUCAAAAGUGGUAGGGUCAUUAUGCUAACGGCCAAUC .((((....................((((((.......((....))....)))))).............(((((((((((((......)))))))))))))............))))... (-24.27 = -25.27 + 1.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:31:22 2006