
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,139,003 – 12,139,143 |
| Length | 140 |
| Max. P | 0.973112 |

| Location | 12,139,003 – 12,139,116 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 82.39 |
| Mean single sequence MFE | -31.48 |
| Consensus MFE | -22.94 |
| Energy contribution | -22.87 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.973112 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
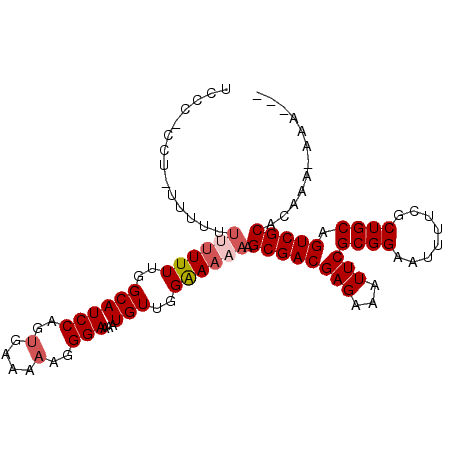
>X_DroMel_CAF1 12139003 113 + 22224390 ACUCCCGUUUUUUUCUCUCGCGCGAUUUCUGUUCAUUUUUUCUC-CUCCUCUUCUUUUUUUGGCAUCCAGUGAAAAAGGGAAAAUGUUGGAAAAAAGCGACGAGAAAUUCGCGG ....((((...((((((((((..((.......)).((((((((.-......((((((((((.((.....)).))))))))))......)))))))))))).))))))...)))) ( -31.72) >DroSec_CAF1 12187 111 + 1 AUG-CCGU-UUUUUCUCUCGCGCGAUUUCUGUUCAUUUUUUCCU-CCUCUUUCUUUUUUUUGGCAUCCAGUGAAAAAGGGAAAAUGUUGGAAAAAAGCGACGAGAAAUUCGCGG ...-((((-..((((((((((..((.......)).((((((((.-.(..(((((((((((((........)))))))))))))..)..)))))))))))).))))))...)))) ( -33.90) >DroEre_CAF1 11839 98 + 1 AUC-CUGU-UUUUUCUCUCGCGCGAUUUGUA----UUUUUUCCC--CU-U-------UUUUGGCAUCCAGUGAAAAAGGGAGAAUGUGGGAAAAAAGCGACGAGAAAUUCGCGG ...-((((-..((((((((((....(((.((----(.(((((((--..-(-------((((.((.....)).)))))))))))).))).)))....)))).))))))...)))) ( -30.80) >DroYak_CAF1 8684 107 + 1 AUU-CGGU-UUUUUCUCUCGAGCGAUUUGUA----CUUUUUUCCGCCU-UUUUUUGUUUUUGGCAUCCAGCGAGAAAGGGAAAAUGUUGGGAAAAAGCGACGAGAAAUUCGCGG ...-(((.-..(((((((((.((.....)).----((((((((((.(.-((((((.(((((.((.....)).))))).)))))).).))))))))))))).)))))).)))... ( -29.50) >consensus AUU_CCGU_UUUUUCUCUCGCGCGAUUUCUA____UUUUUUCCC_CCU_UUUUUUUUUUUUGGCAUCCAGUGAAAAAGGGAAAAUGUUGGAAAAAAGCGACGAGAAAUUCGCGG ....((((...((((((((((....((((((......(((((((............(((((.((.....)).))))))))))))...))))))...)))).))))))...)))) (-22.94 = -22.87 + -0.06)



| Location | 12,139,043 – 12,139,143 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 86.68 |
| Mean single sequence MFE | -26.35 |
| Consensus MFE | -19.19 |
| Energy contribution | -20.00 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.799637 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12139043 100 + 22224390 UCUC-CUCCUCUUCUUUUUUUGGCAUCCAGUGAAAAAGGGAAAAUGUUGGAAAAAAGCGACGAGAAAUUCGCGGAAUUUUCGCUGCAGUCGCACAAAAAAA--- ..((-(..(..((((((((((.((.....)).))))))))))...)..))).....((((((((...)))((((........)))).))))).........--- ( -25.60) >DroSec_CAF1 12225 99 + 1 UCCU-CCUCUUUCUUUUUUUUGGCAUCCAGUGAAAAAGGGAAAAUGUUGGAAAAAAGCGACGAGAAAUUCGCGGAAUUUUCGCUGCAGUCGCACAAA-AAA--- ....-........(((((((..((((((..(....)..)))...)))..)))))))((((((((...)))((((........)))).))))).....-...--- ( -25.70) >DroEre_CAF1 11873 90 + 1 UCCC--CU-U-------UUUUGGCAUCCAGUGAAAAAGGGAGAAUGUGGGAAAAAAGCGACGAGAAAUUCGCGGAAUUUUCGCUGCAGUCGCACAAA-AAA--- ((.(--((-(-------((((.((.....)).)))))))).)).((((.((....(((((....((((((...)))))))))))....)).))))..-...--- ( -26.60) >DroYak_CAF1 8718 102 + 1 UUCCGCCU-UUUUUUGUUUUUGGCAUCCAGCGAGAAAGGGAAAAUGUUGGGAAAAAGCGACGAGAAAUUCGCGGAAUUUUCGCUGCAGUCGCACAAA-AAAAUG (((((.(.-((((((.(((((.((.....)).))))).)))))).).)))))....((((((((...)))((((........)))).))))).....-...... ( -27.50) >consensus UCCC_CCU_UUUUUUUUUUUUGGCAUCCAGUGAAAAAGGGAAAAUGUUGGAAAAAAGCGACGAGAAAUUCGCGGAAUUUUCGCUGCAGUCGCACAAA_AAA___ ..............((((((..((((((..........)))...)))..)))))).((((((((...)))((((........)))).)))))............ (-19.19 = -20.00 + 0.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:30:03 2006