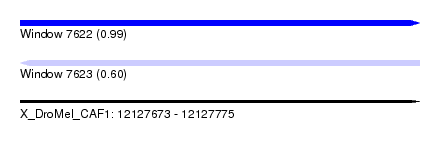

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,127,673 – 12,127,775 |
| Length | 102 |
| Max. P | 0.994817 |

| Location | 12,127,673 – 12,127,775 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 92.94 |
| Mean single sequence MFE | -25.16 |
| Consensus MFE | -23.85 |
| Energy contribution | -23.72 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.95 |
| SVM decision value | 2.51 |
| SVM RNA-class probability | 0.994817 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12127673 102 + 22224390 GCCGCGCAGAGGGCGAAAAAGCCAAACAGGGAGGUUCCUGUAAAUGGUUGGUAAAAAGCCAAUAGCAGACAUUGCUGUGUUAAAUACAAAGAAAGAAAGAAA ...((((((..(((......)))..((((((....)))))).....((((((.....))))))...........))))))...................... ( -24.10) >DroSec_CAF1 2407 101 + 1 GCUGCGCAGAGGGCGAAAAAGCCAAACAGGGAGGUUCCUGUAAAUGGUUGGUAAAAAGCCAAUGGCAGACAUUGCUGUGUUAAAUACAA-UAAAGAAAGAAA .((((.((...(((.....(((((.((((((....))))))...)))))........)))..))))))...(((.(((((...))))).-)))......... ( -25.22) >DroSim_CAF1 2384 101 + 1 GCUGCGCAGAGGGCGAAAAAGCCAAACAGGGAGGUUCCUGUAAAUGGUUGGUAAAAAGCCAAUGGCAGACAUUGCUGUGUUAAAUACAA-UAAAGAAAGAAA .((((.((...(((.....(((((.((((((....))))))...)))))........)))..))))))...(((.(((((...))))).-)))......... ( -25.22) >DroEre_CAF1 2091 98 + 1 GCUUCGCAGGGGGCGAAAAAGCCAAACAGGGAGGGUCCUGUAAAUUAUUGGUAAAAAGCCAAUAGCAGACACUGCUGCGUUAAAUACAA-UAAAGAA---AA ((...((((..(((......)))..((((((....))))))...((((((((.....))))))))......)))).))...........-.......---.. ( -26.10) >consensus GCUGCGCAGAGGGCGAAAAAGCCAAACAGGGAGGUUCCUGUAAAUGGUUGGUAAAAAGCCAAUAGCAGACAUUGCUGUGUUAAAUACAA_UAAAGAAAGAAA ...((((((..(((......)))..((((((....)))))).....((((((.....))))))...........))))))...................... (-23.85 = -23.72 + -0.13)


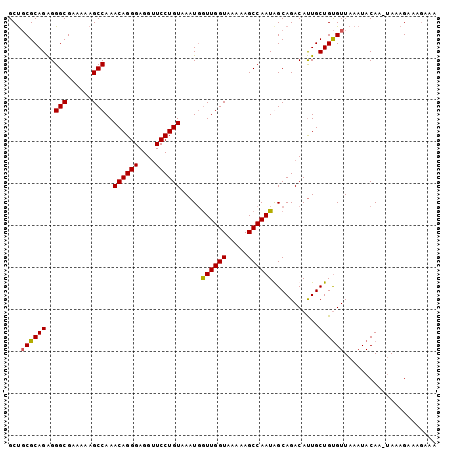
| Location | 12,127,673 – 12,127,775 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 92.94 |
| Mean single sequence MFE | -19.14 |
| Consensus MFE | -15.17 |
| Energy contribution | -14.79 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.600211 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12127673 102 - 22224390 UUUCUUUCUUUCUUUGUAUUUAACACAGCAAUGUCUGCUAUUGGCUUUUUACCAACCAUUUACAGGAACCUCCCUGUUUGGCUUUUUCGCCCUCUGCGCGGC ........................(((....)))((((..((((.......))))(((...(((((......))))).))).......((.....)))))). ( -16.90) >DroSec_CAF1 2407 101 - 1 UUUCUUUCUUUA-UUGUAUUUAACACAGCAAUGUCUGCCAUUGGCUUUUUACCAACCAUUUACAGGAACCUCCCUGUUUGGCUUUUUCGCCCUCUGCGCAGC ..........((-((((..........)))))).((((((..(((..........(((...(((((......))))).))).......)))...)).)))). ( -19.23) >DroSim_CAF1 2384 101 - 1 UUUCUUUCUUUA-UUGUAUUUAACACAGCAAUGUCUGCCAUUGGCUUUUUACCAACCAUUUACAGGAACCUCCCUGUUUGGCUUUUUCGCCCUCUGCGCAGC ..........((-((((..........)))))).((((((..(((..........(((...(((((......))))).))).......)))...)).)))). ( -19.23) >DroEre_CAF1 2091 98 - 1 UU---UUCUUUA-UUGUAUUUAACGCAGCAGUGUCUGCUAUUGGCUUUUUACCAAUAAUUUACAGGACCCUCCCUGUUUGGCUUUUUCGCCCCCUGCGAAGC ..---.......-..........(((((..(((...((((((((.......))))).....(((((......)))))..))).....)))...))))).... ( -21.20) >consensus UUUCUUUCUUUA_UUGUAUUUAACACAGCAAUGUCUGCCAUUGGCUUUUUACCAACCAUUUACAGGAACCUCCCUGUUUGGCUUUUUCGCCCUCUGCGCAGC ........................(((....)))((((((..(((......((((......(((((......))))))))).......)))...)).)))). (-15.17 = -14.79 + -0.38)


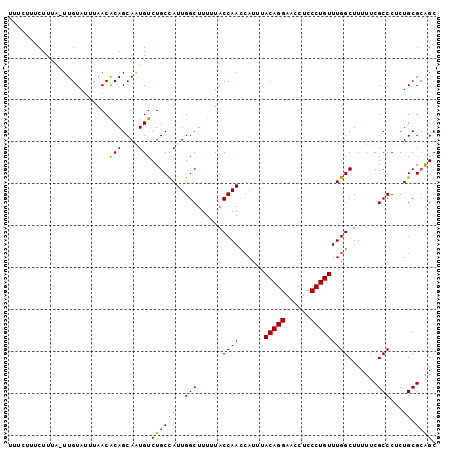
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:29:59 2006