| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,097,173 – 12,097,272 |
| Length | 99 |
| Max. P | 0.932223 |

| Location | 12,097,173 – 12,097,272 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 83.77 |
| Mean single sequence MFE | -31.30 |
| Consensus MFE | -19.19 |
| Energy contribution | -19.31 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.28 |
| Structure conservation index | 0.61 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924367 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12097173 99 + 22224390 AGCUCCCAACGCGCUG---CCUGUUUGCCUUGGCAUUUCUCAAACGGAGAAAUAAAACAAAAAAAAAAAGCAAAAGGAGAUGGUGGCGAC---GAAAAGCGAAGG .(((.....(((((..---((.((((.((((.(((((((((.....)))))))................))..))))))))))..))).)---)...)))..... ( -25.29) >DroEre_CAF1 21174 95 + 1 AGUUCCCAACGCGCUGCCGCCUGUUUGCCUUGGCAUUUCUCAAAU-GAGAAAUGA---------AAAAAGCAAAAGGAGAUGGUGGCGGCGUUGAAAAGCGAAGG .(((..((((((((..((.(((.(((((.((..((((((((....-)))))))).---------..)).))))))))....))..)).))))))...)))..... ( -37.40) >DroYak_CAF1 21613 92 + 1 AGUUCCCAACGCGCUG---CCUGUUUGCCUUGGCAUUUCUCAAAU-GAGAAAUGA---------AAAAAGCAAAAGGAGAUGGCGGUGGCGGUAAAAAGCGGAGG ..((((.....(((((---((..(((((.((..((((((((....-)))))))).---------..)).))))).......)))))))((........)))))). ( -31.20) >consensus AGUUCCCAACGCGCUG___CCUGUUUGCCUUGGCAUUUCUCAAAU_GAGAAAUGA_________AAAAAGCAAAAGGAGAUGGUGGCGGCG_UGAAAAGCGAAGG .....((..(((((((...(((.(((((.((..((((((((.....))))))))............)).))))))))...))))(....)........)))..)) (-19.19 = -19.31 + 0.11)


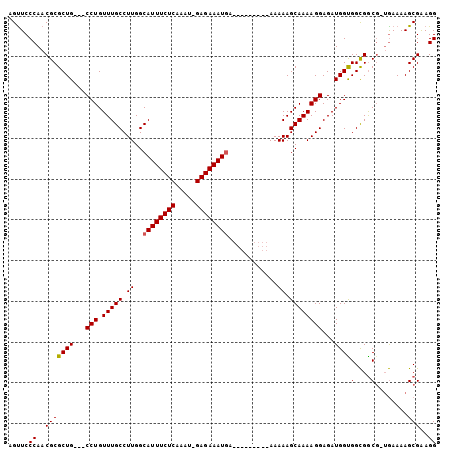
| Location | 12,097,173 – 12,097,272 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 83.77 |
| Mean single sequence MFE | -28.30 |
| Consensus MFE | -17.61 |
| Energy contribution | -17.51 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.25 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12097173 99 - 22224390 CCUUCGCUUUUC---GUCGCCACCAUCUCCUUUUGCUUUUUUUUUUUGUUUUAUUUCUCCGUUUGAGAAAUGCCAAGGCAAACAGG---CAGCGCGUUGGGAGCU .....(((((.(---(.(((.............(((((.(((.(((((...((((((((.....)))))))).))))).))).)))---)))))))...))))). ( -21.51) >DroEre_CAF1 21174 95 - 1 CCUUCGCUUUUCAACGCCGCCACCAUCUCCUUUUGCUUUUU---------UCAUUUCUC-AUUUGAGAAAUGCCAAGGCAAACAGGCGGCAGCGCGUUGGGAACU .......((..((((((.((..((....(((((((((((..---------.((((((((-....))))))))..)))))))).))).))..))))))))..)).. ( -35.00) >DroYak_CAF1 21613 92 - 1 CCUCCGCUUUUUACCGCCACCGCCAUCUCCUUUUGCUUUUU---------UCAUUUCUC-AUUUGAGAAAUGCCAAGGCAAACAGG---CAGCGCGUUGGGAACU ..(((((........))((.(((...(((((((((((((..---------.((((((((-....))))))))..)))))))).)))---.)).))).)))))... ( -28.40) >consensus CCUUCGCUUUUCA_CGCCGCCACCAUCUCCUUUUGCUUUUU_________UCAUUUCUC_AUUUGAGAAAUGCCAAGGCAAACAGG___CAGCGCGUUGGGAACU ((..(((........((....)).....(((((((((((............((((((((.....))))))))..)))))))).))).......)))..))..... (-17.61 = -17.51 + -0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:29:41 2006