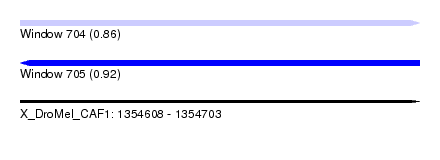

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,354,608 – 1,354,703 |
| Length | 95 |
| Max. P | 0.916444 |

| Location | 1,354,608 – 1,354,703 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 95.57 |
| Mean single sequence MFE | -37.84 |
| Consensus MFE | -36.84 |
| Energy contribution | -36.48 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.859301 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1354608 95 + 22224390 GGAAAAAGGUCCGGGGCGUGGCAGGAUUAGGCCCGAGGGCCAUUCUUGACCUGCUAGUGACUCCAAGGAUGCGCCUUCUUCUUACGACGGCGCCA ........(((((((((((((((((.(((((((....)))).....)))))))))).)).))))..))))(((((.((.......)).))))).. ( -38.00) >DroSec_CAF1 14828 94 + 1 GGAAAAA-GUCUGGGGCGUGGCAGGAUUAGGCCCGAGGGCCAUUCUUGACCUGCUAGUGACUCCAAGGAUGCGCCUUCUUCUUACGACGGCGCCA (((....-(((...((((.(((((((...((((....))))..))))).))))))...))))))......(((((.((.......)).))))).. ( -37.40) >DroSim_CAF1 18280 94 + 1 GGAAAAA-GUCUGGGGCGUGGCAGGAUUAGGCCCGAGGGCCAUUCUUGACCUGCUAGUGACUCCAAGGAUGCGCCUUCUUCUUACGACGGCGCCA (((....-(((...((((.(((((((...((((....))))..))))).))))))...))))))......(((((.((.......)).))))).. ( -37.40) >DroEre_CAF1 15156 95 + 1 GGGAAAAGGUCUGGGGCGUGGCAGGAUUAGGCUCGAGGGCCAUUCUUGACCUGCCAGUGACUCCAAGGAUGCGCCUCCUUCUUGCGACGGCGCCA .(((....(((...((((.(((((((...((((....))))..))))).))))))...))))))......((((((((.....).)).))))).. ( -36.70) >DroYak_CAF1 15565 95 + 1 GGAAAGCGGUCUGGGGCGUGGCAGGAUUAGGCCCGAGGGCCAUUCUUGACCUGCCAGUGACUCCAAGGAUGCGCCUUCUUCUUGCGACGGCGCCA ((...((.(((.(((((((((((((.(((((((....)))).....)))))))))).)).))))((((.....))))........))).)).)). ( -39.70) >consensus GGAAAAAGGUCUGGGGCGUGGCAGGAUUAGGCCCGAGGGCCAUUCUUGACCUGCUAGUGACUCCAAGGAUGCGCCUUCUUCUUACGACGGCGCCA (((.....(((...((((.(((((((...((((....))))..))))).))))))...))))))......(((((.((.......)).))))).. (-36.84 = -36.48 + -0.36)

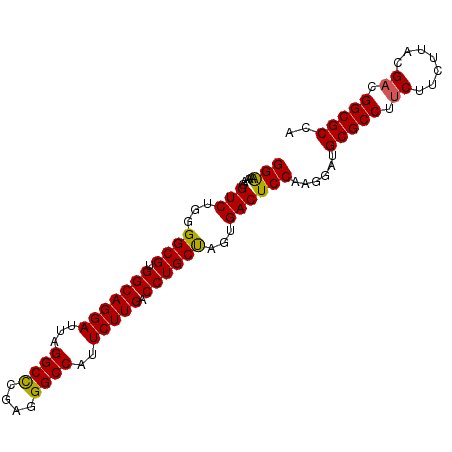
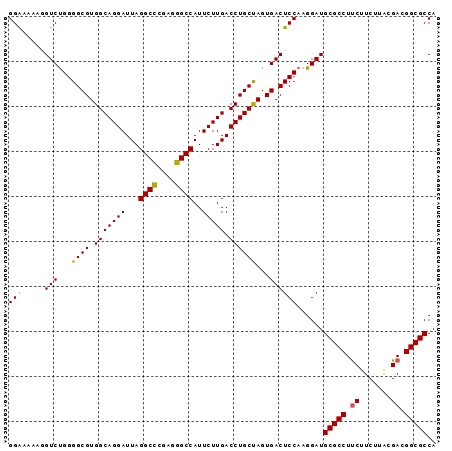
| Location | 1,354,608 – 1,354,703 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 95.57 |
| Mean single sequence MFE | -34.60 |
| Consensus MFE | -31.86 |
| Energy contribution | -31.90 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.916444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1354608 95 - 22224390 UGGCGCCGUCGUAAGAAGAAGGCGCAUCCUUGGAGUCACUAGCAGGUCAAGAAUGGCCCUCGGGCCUAAUCCUGCCACGCCCCGGACCUUUUUCC ..(((((.((....))....))))).(((..((.((.....(((((........((((....))))....))))).)).))..)))......... ( -33.80) >DroSec_CAF1 14828 94 - 1 UGGCGCCGUCGUAAGAAGAAGGCGCAUCCUUGGAGUCACUAGCAGGUCAAGAAUGGCCCUCGGGCCUAAUCCUGCCACGCCCCAGAC-UUUUUCC ..(((((.((....))....)))))....((((.((.....(((((........((((....))))....)))))...)).))))..-....... ( -32.90) >DroSim_CAF1 18280 94 - 1 UGGCGCCGUCGUAAGAAGAAGGCGCAUCCUUGGAGUCACUAGCAGGUCAAGAAUGGCCCUCGGGCCUAAUCCUGCCACGCCCCAGAC-UUUUUCC ..(((((.((....))....)))))....((((.((.....(((((........((((....))))....)))))...)).))))..-....... ( -32.90) >DroEre_CAF1 15156 95 - 1 UGGCGCCGUCGCAAGAAGGAGGCGCAUCCUUGGAGUCACUGGCAGGUCAAGAAUGGCCCUCGAGCCUAAUCCUGCCACGCCCCAGACCUUUUCCC ..(((((.((....))....)))))....((((.((...(((((((........(((......)))....))))))).)).)))).......... ( -34.20) >DroYak_CAF1 15565 95 - 1 UGGCGCCGUCGCAAGAAGAAGGCGCAUCCUUGGAGUCACUGGCAGGUCAAGAAUGGCCCUCGGGCCUAAUCCUGCCACGCCCCAGACCGCUUUCC ..(((((.((....))....)))))....((((.((...(((((((........((((....))))....))))))).)).)))).......... ( -39.20) >consensus UGGCGCCGUCGUAAGAAGAAGGCGCAUCCUUGGAGUCACUAGCAGGUCAAGAAUGGCCCUCGGGCCUAAUCCUGCCACGCCCCAGACCUUUUUCC ..(((((.((....))....)))))....((((.((.....(((((........((((....))))....)))))...)).)))).......... (-31.86 = -31.90 + 0.04)

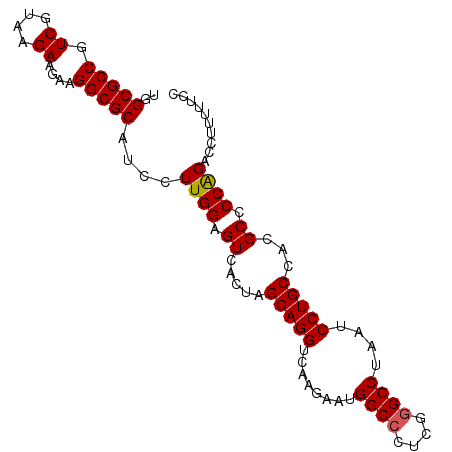
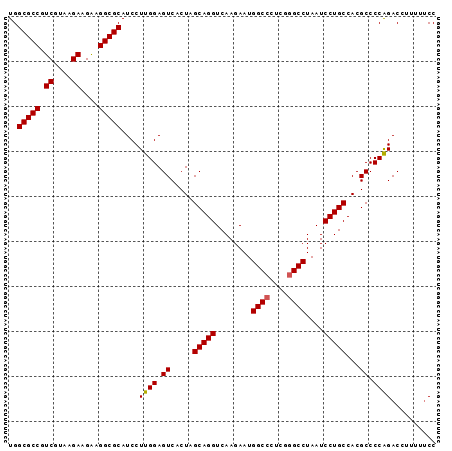
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:45:04 2006