
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,011,041 – 12,011,149 |
| Length | 108 |
| Max. P | 0.897010 |

| Location | 12,011,041 – 12,011,149 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.28 |
| Mean single sequence MFE | -44.83 |
| Consensus MFE | -32.06 |
| Energy contribution | -33.40 |
| Covariance contribution | 1.34 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.897010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12011041 108 + 22224390 GGACUCGGCA------------GCAGCCGAUUCCUGAAUCUUAGUGGUACCAAAUCUACAGCAUUUAGUGAGGGCAACCAACGCCACAAUGGCCACCAGGCCAAUGGCCCAAAUCGAGGG (((.(((((.------------...))))).)))....((((.((((........))))..........((((....))...((((...(((((....))))).)))).....)))))). ( -35.50) >DroSec_CAF1 44569 120 + 1 GGCCUCGGCAUCAGCCUUUUUGGCAUCCGAGUCCUGAACCUUAGCGGUACCAGAUCCACACCAGAUGGUGAGGGCAACCAGCGCCACAAUGGCCAGCAGGCCAAUGGCCGGGCUCGAGGG ..((((.......(((.....)))....((((((.........((((........)).((((....)))).((....)).))((((...(((((....))))).)))).)))))))))). ( -49.50) >DroSim_CAF1 43792 120 + 1 GGCCUCGGCAUUAUCCUUUUUGGCAUCCAAGUACUGAACCUUAUAGGUAUCAGAUCCACACCAGAUGGUGAGGGCAACCAGCGCCACAAUGGCCAGCAGGCCAAUGGCCGGGCCCGAGGG ((((.((((...........((((.........(((((((.....))).)))).....((((....)))).((....))...))))((.(((((....))))).))))))))))...... ( -49.50) >consensus GGCCUCGGCAU_A_CCUUUUUGGCAUCCGAGUCCUGAACCUUAGAGGUACCAGAUCCACACCAGAUGGUGAGGGCAACCAGCGCCACAAUGGCCAGCAGGCCAAUGGCCGGGCUCGAGGG ((.((((((...........((((.............(((.....)))..........((((....)))).((....))...))))((.(((((....))))).)))))))).))..... (-32.06 = -33.40 + 1.34)


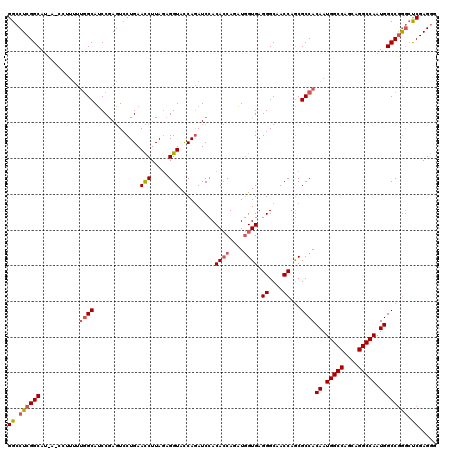
| Location | 12,011,041 – 12,011,149 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.28 |
| Mean single sequence MFE | -47.23 |
| Consensus MFE | -33.35 |
| Energy contribution | -33.47 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.775459 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 12011041 108 - 22224390 CCCUCGAUUUGGGCCAUUGGCCUGGUGGCCAUUGUGGCGUUGGUUGCCCUCACUAAAUGCUGUAGAUUUGGUACCACUAAGAUUCAGGAAUCGGCUGC------------UGCCGAGUCC .......(((((((((((((((....)))))..)))))..((((.(((.((((........)).))...)))))))))))).....(((.(((((...------------.))))).))) ( -37.80) >DroSec_CAF1 44569 120 - 1 CCCUCGAGCCCGGCCAUUGGCCUGCUGGCCAUUGUGGCGCUGGUUGCCCUCACCAUCUGGUGUGGAUCUGGUACCGCUAAGGUUCAGGACUCGGAUGCCAAAAAGGCUGAUGCCGAGGCC .(((((.((.(((((..(((((....)))))...(((((((((((..((.((((....)))).))..((((.(((.....)))))))))).))).)))))....)))))..))))))).. ( -55.30) >DroSim_CAF1 43792 120 - 1 CCCUCGGGCCCGGCCAUUGGCCUGCUGGCCAUUGUGGCGCUGGUUGCCCUCACCAUCUGGUGUGGAUCUGAUACCUAUAAGGUUCAGUACUUGGAUGCCAAAAAGGAUAAUGCCGAGGCC ......((((((((((.(((((....))))).))((((((..(.(((((.((((....)))).))...(((.(((.....))))))))).)..).)))))...........)))).)))) ( -48.60) >consensus CCCUCGAGCCCGGCCAUUGGCCUGCUGGCCAUUGUGGCGCUGGUUGCCCUCACCAUCUGGUGUGGAUCUGGUACCACUAAGGUUCAGGACUCGGAUGCCAAAAAGG_U_AUGCCGAGGCC .(((.(((((..((((((((((....)))))..)))))...(((.(((((((((....)))).))....)))))).....)))))))).(((((..................)))))... (-33.35 = -33.47 + 0.12)

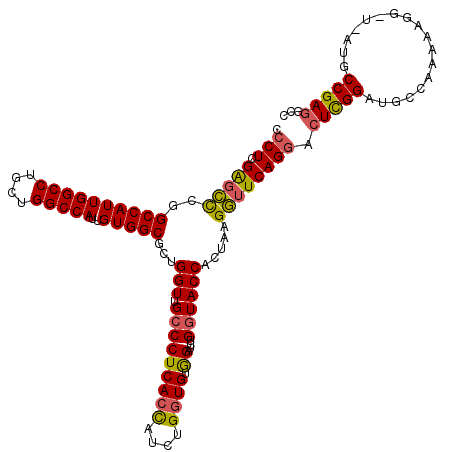

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:28:37 2006