
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,885,853 – 11,885,984 |
| Length | 131 |
| Max. P | 0.891303 |

| Location | 11,885,853 – 11,885,945 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 95.76 |
| Mean single sequence MFE | -27.74 |
| Consensus MFE | -22.02 |
| Energy contribution | -23.26 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.720409 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11885853 92 + 22224390 ACCAUCCAUGCCCUGGGUUCCAAGGAUCUGGGCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAA .(((...((((((.(((((.....)))))))))))......)))((((.((((..(((((((((((....)))))).))))).)))))))). ( -30.40) >DroSec_CAF1 17762 92 + 1 ACCAACCAUGUCCUGGGUUCCAAGGAUCUGGGCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAA .((((..(((.((..((((.....))))..))))).....))))((((.((((..(((((((((((....)))))).))))).)))))))). ( -28.20) >DroSim_CAF1 22341 92 + 1 ACCACCCAUGUCCUGGGUUCCAAGGAUCUGGGCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAA .(((((((.(((((.(....).))))).)))).........)))((((.((((..(((((((((((....)))))).))))).)))))))). ( -28.90) >DroEre_CAF1 19285 91 + 1 ACCACCCAUGUCCUGGGCUCCAAGGAUCUG-GCAUUCGCUUGGGUCUCUGUUUGCCGUGGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAA ....((((.(((((.(....).)))))..(-((....)))))))((((.((((..(((((((((((....)))))))).))).)))))))). ( -27.00) >DroYak_CAF1 21967 91 + 1 ACCACCCAUGUCCUGGGCUCCAAGGAUCUG-GCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGACAUAAUCAACGAAAACGAGAA .....(((.(((((.(....).))))).))-)............((((.((((..(((((.((((....))))....))))).)))))))). ( -24.20) >consensus ACCACCCAUGUCCUGGGUUCCAAGGAUCUGGGCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAA .(((...((((((.(((((.....)))))))))))......)))((((.((((..(((((((((((....)))))).))))).)))))))). (-22.02 = -23.26 + 1.24)
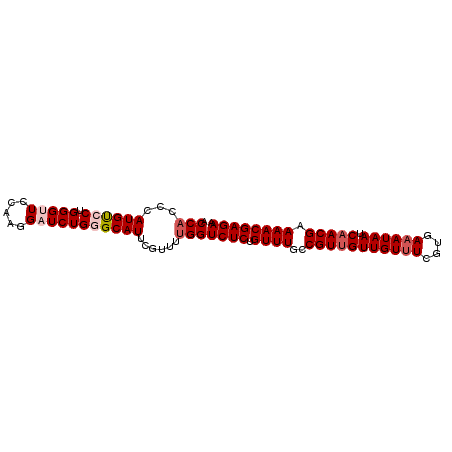



| Location | 11,885,884 – 11,885,984 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 94.41 |
| Mean single sequence MFE | -19.82 |
| Consensus MFE | -15.80 |
| Energy contribution | -16.04 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.891303 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11885884 100 + 22224390 GCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAAAA-AAAAUUGAAAGUAAGCAAGAAAUAAAUAAUAAAACAA ((.((((((((..((((.((((..(((((((((((....)))))).))))).))))))))...-)))).)))).))......................... ( -19.90) >DroSec_CAF1 17793 101 + 1 GCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAAAAAAAAAUUGAAAGUAAGCAAGAAAUAAAUAAUAAAACAA ((.((((((((..((((.((((..(((((((((((....)))))).))))).))))))))....)))).)))).))......................... ( -19.20) >DroSim_CAF1 22372 99 + 1 GCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAAA--AAAAUUGAAAGUAAGCAAGAAAUAAAUAAUAAAACAA ((.((((((((..((((.((((..(((((((((((....)))))).))))).))))))))..--)))).)))).))......................... ( -20.90) >DroEre_CAF1 19315 97 + 1 GCAUUCGCUUGGGUCUCUGUUUGCCGUGGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAAAA---AA-UGAGAGUAGGCUAGAAAUAAAUAAUAAAACAA ((...(.((((..((((.((((..(((((((((((....)))))))).))).))))))))...---..-)))).)...))..................... ( -19.30) >DroYak_CAF1 21997 98 + 1 GCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGACAUAAUCAACGAAAACGAGAAAA---AAUUGAAAGUAAGCAAAAAAUAAAUAAUAAAACAA ((.((((.(((..((((.((((..(((((.((((....))))....))))).))))))))..)---)).)))).))......................... ( -19.80) >consensus GCAUUCGUUUUGGUCUCUGUUUGCCGUUGUUGUUUCGUGAAAUAAUCAACGAAAACGAGAAAA_AAAAUUGAAAGUAAGCAAGAAAUAAAUAAUAAAACAA ((.((((......((((.((((..(((((((((((....)))))).))))).)))))))).........)))).))......................... (-15.80 = -16.04 + 0.24)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:26:40 2006