
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,824,429 – 11,824,596 |
| Length | 167 |
| Max. P | 0.765970 |

| Location | 11,824,429 – 11,824,543 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.17 |
| Mean single sequence MFE | -52.93 |
| Consensus MFE | -28.71 |
| Energy contribution | -30.13 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.587427 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11824429 114 - 22224390 CAGCUGCAGCUGCAGCGGCGGCUCAUCCUUCGGAGGCUCCUGCUCCAGUCG---AAGCAGAAGCUCACGCCGCAGAGGCUCCCGCUGCCGUGGAAGGAGAUGCC---GCACCAGCGGCUG (((((((.((....)).(((((...((((((.(((.((.(((((.(....)---.))))).))))).(((.((((.((...)).)))).)))))))))...)))---))....))))))) ( -56.60) >DroSec_CAF1 237 114 - 1 CAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCCUGCUCCAGUCG---AAGCAGAAGCUACCGCCGCAGAGGCUCCCGCUGCCGUGGAAGGAGAUGCC---GCACCAGCGGCUG (((((((.((....)).(((((...((((((((.((((.(((((.(....)---.))))).)))).))((.((((.((...)).)))).)))).))))...)))---))....))))))) ( -56.40) >DroSim_CAF1 159 114 - 1 CAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCCUGCUCCAGUCG---AAGGCGAAGCUCCCGCCGCAGAGGCUCCCGCUGCCGUGGAAGGAGAUGCC---GCACCAGCGGCUG (((((((.((....)).(((((...((((((((..((..((((((.....)---).((((.......))))))))..))..))((....)))).))))...)))---))....))))))) ( -53.20) >DroEre_CAF1 1755 111 - 1 C------AGCUGCAGCGGCGGCUCAUCCACCGGAGGCACAUGCUCCAGCCGCAGAGGCAGAAGCGCCCGCCGCGGAGGCUCCCGCUGGCGAAGAAGGAGCUGUC---GCACCAGCGGCCG .------.(((((.((((((((((.(((...)))(((....(((.(.(((.....))).).))))))((((((((......)))).))))......))))))))---))....))))).. ( -51.50) >DroYak_CAF1 231 108 - 1 C------AGCUGCAGCGGCGGCUCAUCCUCCGGAGCCACCUGCUCCAGUCG---AAGCCGAAGCUCCCGCCGCUGAAGCUCCCGCUGGCGACGAAGGAGCUGCC---GCACCAGCAGCUG (------((((((.(((((((((..((..(.(((((.....))))).)..)---)))))..((((((((.((((..(((....))))))).))..)))))))))---))....))))))) ( -55.40) >DroAna_CAF1 10465 96 - 1 C---GGCGGCUGCGGCUGC------------GA---AUCCGGCUUCAGUAG---AGGCG---GCUCCGGCAGAAGGGGCGGCGCCGGCAGGAGAAGGAGAAGCCCCUGCAACCGUUGUUG (---(((((.(((((..((------------..---.(((..((((.((..---.((((---(((((.......)))))..)))).)).))))..)))...))..))))).))))))... ( -44.50) >consensus C___UGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCCUGCUCCAGUCG___AAGCAGAAGCUCCCGCCGCAGAGGCUCCCGCUGCCGUGGAAGGAGAUGCC___GCACCAGCGGCUG ......(((((((.(((((((..........((((.......)))).........(((....))).))))))).....((((..((.....))..))))..............))))))) (-28.71 = -30.13 + 1.42)



| Location | 11,824,466 – 11,824,571 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 85.70 |
| Mean single sequence MFE | -49.34 |
| Consensus MFE | -32.84 |
| Energy contribution | -36.44 |
| Covariance contribution | 3.60 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.749712 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11824466 105 + 22224390 AGCCUCUGCGGCGUGAGCUUCUGCUU---CGACUGGAGCAGGAGCCUCCGAAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCCGCUGCCUCCAUUUCCUGAUUGAU .(((.....)))(.((((((((((((---(....)))))))))).))))..((((.((((.((.((((((((....)))))))).)).).)))....))))....... ( -49.30) >DroSec_CAF1 274 105 + 1 AGCCUCUGCGGCGGUAGCUUCUGCUU---CGACUGGAGCAGGAGCCUCCGGAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCCGCAGCCUCCAUUUCCUGAUUGAU .(((.....)))((..((((((((((---(....)))))))))))..))((((((.(.((.((.((((((((....)))))))).)).))))))...)))........ ( -53.90) >DroSim_CAF1 196 105 + 1 AGCCUCUGCGGCGGGAGCUUCGCCUU---CGACUGGAGCAGGAGCCUCCGGAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCUGCAGCCUCCACUUCCUGAUUGAU .(((.....)))((..(((((..(((---(....))))..)))))..))(((((..(((.....(((((((((((....))))))))))))))..)))))........ ( -49.60) >DroEre_CAF1 1792 102 + 1 AGCCUCCGCGGCGGGCGCUUCUGCCUCUGCGGCUGGAGCAUGUGCCUCCGGUGGAUGAGCCGCCGCUGCAGCU------GCUGCCGCCGUCUCCAUUUCCUGAGUGAU .......(((((((..((..((((....(((((.((..(((.((((...)))).)))..))))))).))))..------))..)))))))...(((((...))))).. ( -43.30) >DroYak_CAF1 268 99 + 1 AGCUUCAGCGGCGGGAGCUUCGGCUU---CGACUGGAGCAGGUGGCUCCGGAGGAUGAGCCGCCGCUGCAGCU------GCUGCCGCUGUCUCCACUUCCUGGGUGAU .....(((((((((((((....))))---.(.(((.(((.((((((((........))))))))))).)))).------.)))))))))....((((.....)))).. ( -50.60) >consensus AGCCUCUGCGGCGGGAGCUUCUGCUU___CGACUGGAGCAGGAGCCUCCGGAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCCGCAGCCUCCAUUUCCUGAUUGAU .(((.....)))(((.((((((((((.........)))))))))).)))(((((..((((.((.((((((((....)))))))).)).).)))..)))))........ (-32.84 = -36.44 + 3.60)



| Location | 11,824,466 – 11,824,571 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 85.70 |
| Mean single sequence MFE | -46.90 |
| Consensus MFE | -31.68 |
| Energy contribution | -33.12 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751095 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11824466 105 - 22224390 AUCAAUCAGGAAAUGGAGGCAGCGGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUUCGGAGGCUCCUGCUCCAGUCG---AAGCAGAAGCUCACGCCGCAGAGGCU .......((((....(((.(.((.((((((((....)))))))).))).))).))))..(..((((.(((((.(....)---.))))).))))..)(((.....))). ( -47.60) >DroSec_CAF1 274 105 - 1 AUCAAUCAGGAAAUGGAGGCUGCGGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCCUGCUCCAGUCG---AAGCAGAAGCUACCGCCGCAGAGGCU ........(((...(((((((((.((((((((....)))))))).))))))..))))))((.((((.(((((.(....)---.))))).)))).))(((.....))). ( -54.30) >DroSim_CAF1 196 105 - 1 AUCAAUCAGGAAGUGGAGGCUGCAGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCCUGCUCCAGUCG---AAGGCGAAGCUCCCGCCGCAGAGGCU .....((.(((...(((((((((.((((((((....)))))))).))))))..)))))).))(((..((((((.....)---).((((.......))))))))..))) ( -49.30) >DroEre_CAF1 1792 102 - 1 AUCACUCAGGAAAUGGAGACGGCGGCAGC------AGCUGCAGCGGCGGCUCAUCCACCGGAGGCACAUGCUCCAGCCGCAGAGGCAGAAGCGCCCGCCGCGGAGGCU ....(((.......(....)(((((..((------..((((.(((((((........))((((.(....))))).)))))....))))..))..)))))...)))... ( -42.80) >DroYak_CAF1 268 99 - 1 AUCACCCAGGAAGUGGAGACAGCGGCAGC------AGCUGCAGCGGCGGCUCAUCCUCCGGAGCCACCUGCUCCAGUCG---AAGCCGAAGCUCCCGCCGCUGAAGCU .((((.......))))...(((((((.((------....))(((..(((((..((..(.(((((.....))))).)..)---))))))..)))...)))))))..... ( -40.50) >consensus AUCAAUCAGGAAAUGGAGGCAGCGGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCCUGCUCCAGUCG___AAGCAGAAGCUCCCGCCGCAGAGGCU .((..................((((((((....)))))))).(((((((..........((((.(....)))))..........((....))..))))))).)).... (-31.68 = -33.12 + 1.44)



| Location | 11,824,503 – 11,824,596 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 85.35 |
| Mean single sequence MFE | -39.68 |
| Consensus MFE | -28.98 |
| Energy contribution | -30.78 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.753758 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11824503 93 + 22224390 GGAGCCUCCGAAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCCGCUGCCUCCAUUUCCUGAUUGAUUUCGGUGGCCCGCUUGCUCUGAAGA--- ((.(((.((((((((.(.((.((.((((((((....)))))))).)).))))))...((......)).))))).)))))..............--- ( -40.60) >DroSec_CAF1 311 92 + 1 GGAGCCUCCGGAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCCGCAGCCUCCAUUUCCUGAUUGAUUUCGGUGGCCCGCUUGCUCUGAAA---- ((.(((.((((((((.(.((.((.((((((((....)))))))).)).))))))...((......)).))))).))))).............---- ( -40.10) >DroSim_CAF1 233 96 + 1 GGAGCCUCCGGAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCUGCAGCCUCCACUUCCUGAUUGAUUUCGGUGGCCCGCUUGCUCUGCAGAAAG ((.(((.(((((((..(((.....(((((((((((....))))))))))))))..))))).((......)))).)))))((.......))...... ( -44.30) >DroEre_CAF1 1832 90 + 1 UGUGCCUCCGGUGGAUGAGCCGCCGCUGCAGCU------GCUGCCGCCGUCUCCAUUUCCUGAGUGAUUUCGGUGGCCCGCUUGCUCUGAAAAGAU ..(((...((((((.....))))))..)))((.------((.(((((((....(((((...)))))....)))))))..))..))(((....))). ( -33.00) >DroYak_CAF1 305 89 + 1 GGUGGCUCCGGAGGAUGAGCCGCCGCUGCAGCU------GCUGCCGCUGUCUCCACUUCCUGGGUGAUUUCGGUGGCCCGCUUGCUCUGCAAU-AA ((((((((........))))))))((.(((((.------((..(((..(((((((.....)))).)))..)))..))..)).)))...))...-.. ( -40.40) >consensus GGAGCCUCCGGAGGAUGAGCCGCCGCUGCAGCUGCAGCUGCAGCCGCAGCCUCCAUUUCCUGAUUGAUUUCGGUGGCCCGCUUGCUCUGAAAA_A_ ((.(((.((((((((.(.((.((.((((((((....)))))))).)).))))))...((......)).))))).)))))................. (-28.98 = -30.78 + 1.80)

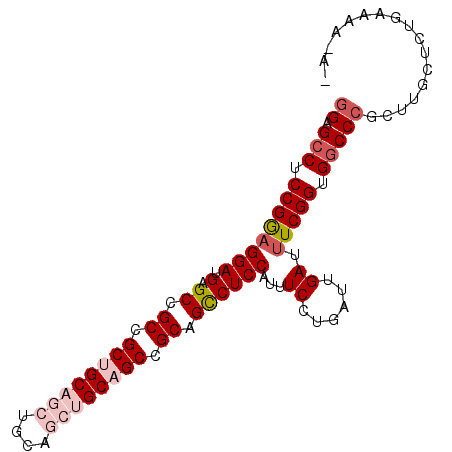

| Location | 11,824,503 – 11,824,596 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 85.35 |
| Mean single sequence MFE | -37.92 |
| Consensus MFE | -29.56 |
| Energy contribution | -31.04 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.765970 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11824503 93 - 22224390 ---UCUUCAGAGCAAGCGGGCCACCGAAAUCAAUCAGGAAAUGGAGGCAGCGGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUUCGGAGGCUCC ---..............(((((.((((........((((....(((.(.((.((((((((....)))))))).))).))).)))))))).))))). ( -43.00) >DroSec_CAF1 311 92 - 1 ----UUUCAGAGCAAGCGGGCCACCGAAAUCAAUCAGGAAAUGGAGGCUGCGGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCC ----.............(((((.((((......)).(((...(((((((((.((((((((....)))))))).))))))..)))))))).))))). ( -45.30) >DroSim_CAF1 233 96 - 1 CUUUCUGCAGAGCAAGCGGGCCACCGAAAUCAAUCAGGAAGUGGAGGCUGCAGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCC ......((.......))(((((.((((......)).(((...(((((((((.((((((((....)))))))).))))))..)))))))).))))). ( -44.20) >DroEre_CAF1 1832 90 - 1 AUCUUUUCAGAGCAAGCGGGCCACCGAAAUCACUCAGGAAAUGGAGACGGCGGCAGC------AGCUGCAGCGGCGGCUCAUCCACCGGAGGCACA ...............(((((((.(((......(((........)))...(((((...------.)))))..))).))))).(((...))).))... ( -27.90) >DroYak_CAF1 305 89 - 1 UU-AUUGCAGAGCAAGCGGGCCACCGAAAUCACCCAGGAAGUGGAGACAGCGGCAGC------AGCUGCAGCGGCGGCUCAUCCUCCGGAGCCACC ..-.((((...))))(((((((.(((...............((....))(((((...------.)))))..))).))))).(((...))))).... ( -29.20) >consensus _U_UUUUCAGAGCAAGCGGGCCACCGAAAUCAAUCAGGAAAUGGAGGCAGCGGCUGCAGCUGCAGCUGCAGCGGCGGCUCAUCCUCCGGAGGCUCC .........((((....(((((.(((.......(((.....))).....((((((((....))))))))..))).))))).(((...))).)))). (-29.56 = -31.04 + 1.48)

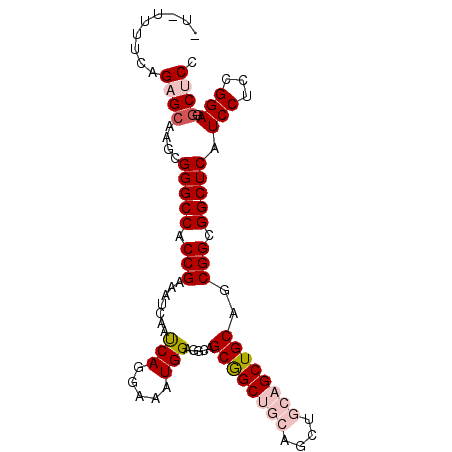

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:25:53 2006