
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,773,832 – 11,773,933 |
| Length | 101 |
| Max. P | 0.933144 |

| Location | 11,773,832 – 11,773,933 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 75.80 |
| Mean single sequence MFE | -25.97 |
| Consensus MFE | -19.74 |
| Energy contribution | -18.97 |
| Covariance contribution | -0.78 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11773832 101 + 22224390 -AUUUUUUUGUUUUUAAC--GUUCUUACCUCAAGCUAGGCGAUCGCCGAUGGCCAUUCAUAUACGAAUGUCCAGGAUGUAAAUGGCCAGCAUCGGCACCGGUAA -.................--.....((((....((...))....(((((((((((((.((((.(.........).)))).))))))))...)))))...)))). ( -25.80) >DroVir_CAF1 4444 78 + 1 --------------------------ACCUCAAGCUAGGCGAAUGCCGAUGGCCAUUUAUAUACGAAUGUCCAGGAUGUAAAUGGCCAGCAUCGGGACCAGUCA --------------------------.(((...(((.(((....)))...((((((((((((.(.........).)))))))))))))))...)))........ ( -26.00) >DroGri_CAF1 3993 98 + 1 UGGUU---UG---UUUGUAAAAAUUAACCUCAAACUAGGCGAAUGCCGAUGGCCAUUCAUAUACGAAUGUCCAGGAUGUAAAUGGCCAGCAUCGGCUCCAGUCA (((((---((---...((........))..)))))))(((....(((((((((((((.((((.(.........).)))).))))))))...)))))....))). ( -30.40) >DroWil_CAF1 7858 87 + 1 -A------------UUGA----ACUCACCUCAAACUGGGCGAACGGCGAUGGCCAUUCAUAUACGAAUGUCCAGGAUGUAAAUGGCCAGCAUCGGCACCGGUAA -.------------((((----.......))))(((((((((...((..((((((((.((((.(.........).)))).)))))))))).)).)).))))).. ( -24.50) >DroYak_CAF1 3715 81 + 1 -----------------------CAUACCUCAGGCUAGGCGAUCGCCGAUGGCCAUUCAUGUACGAAUGUCCAGGAUGUAGAUGGCCAGCAUCGGCACCGGUUA -----------------------..........(((.(((.(((.((..(((.(((((......))))).))))).....))).))))))((((....)))).. ( -25.00) >DroAna_CAF1 3484 91 + 1 -GCCUU-------AAAG-----CCUUACCUUAAACUCGGCGAUUGUCGAUGGCCAUUCAUGUAGGAAAGUCCAGGAUGUAAAUGGCUAGCAUCGGCACCAGUGA -.....-------...(-----((.............)))...(((((((((((((((((.(.((.....)).).)))..))))))...))))))))....... ( -24.12) >consensus ______________U_G______CUUACCUCAAACUAGGCGAACGCCGAUGGCCAUUCAUAUACGAAUGUCCAGGAUGUAAAUGGCCAGCAUCGGCACCAGUAA .................................(((.((.....(((((((((((((.((((..(......)...)))).))))))))...))))).))))).. (-19.74 = -18.97 + -0.78)
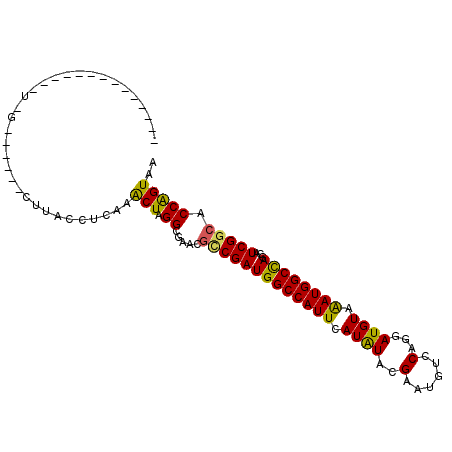


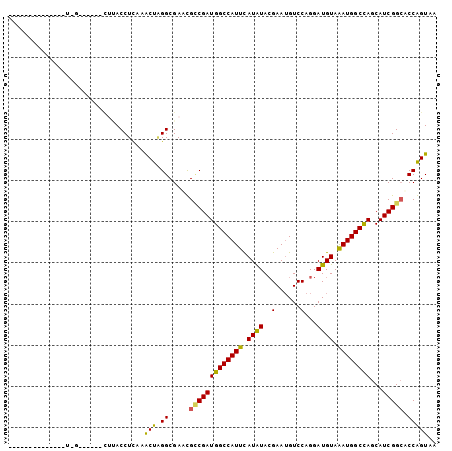
| Location | 11,773,832 – 11,773,933 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 75.80 |
| Mean single sequence MFE | -25.08 |
| Consensus MFE | -17.78 |
| Energy contribution | -18.64 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.12 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11773832 101 - 22224390 UUACCGGUGCCGAUGCUGGCCAUUUACAUCCUGGACAUUCGUAUAUGAAUGGCCAUCGGCGAUCGCCUAGCUUGAGGUAAGAAC--GUUAAAAACAAAAAAAU- (((((((((((((...((((((((((.((.(.........).)).))))))))))))))).)))((...))....)))))....--.................- ( -26.80) >DroVir_CAF1 4444 78 - 1 UGACUGGUCCCGAUGCUGGCCAUUUACAUCCUGGACAUUCGUAUAUAAAUGGCCAUCGGCAUUCGCCUAGCUUGAGGU-------------------------- ........(((((.((((((((((((.((.(.........).)).)))))))))...(((....))).)))))).)).-------------------------- ( -21.20) >DroGri_CAF1 3993 98 - 1 UGACUGGAGCCGAUGCUGGCCAUUUACAUCCUGGACAUUCGUAUAUGAAUGGCCAUCGGCAUUCGCCUAGUUUGAGGUUAAUUUUUACAAA---CA---AACCA .((((((.((.(((((((((((((((.((.(.........).)).)))))))))...)))))).))))))))...((((..(((....)))---..---)))). ( -28.40) >DroWil_CAF1 7858 87 - 1 UUACCGGUGCCGAUGCUGGCCAUUUACAUCCUGGACAUUCGUAUAUGAAUGGCCAUCGCCGUUCGCCCAGUUUGAGGUGAGU----UCAA------------U- (((((.(.((((....))))).....((..((((((....))....(((((((....)))))))..))))..)).)))))..----....------------.- ( -26.30) >DroYak_CAF1 3715 81 - 1 UAACCGGUGCCGAUGCUGGCCAUCUACAUCCUGGACAUUCGUACAUGAAUGGCCAUCGGCGAUCGCCUAGCCUGAGGUAUG----------------------- .....((((((((...(((((((.....((.((.((....)).)).)))))))))))))).)))((((......))))...----------------------- ( -24.80) >DroAna_CAF1 3484 91 - 1 UCACUGGUGCCGAUGCUAGCCAUUUACAUCCUGGACUUUCCUACAUGAAUGGCCAUCGACAAUCGCCGAGUUUAAGGUAAGG-----CUUU-------AAGGC- ...((...(((((((...((((((((......((.....))....)))))))))))).......(((........)))..))-----)...-------.))..- ( -23.00) >consensus UGACCGGUGCCGAUGCUGGCCAUUUACAUCCUGGACAUUCGUAUAUGAAUGGCCAUCGGCAAUCGCCUAGCUUGAGGUAAG______C_A______________ .....((((((((...((((((((((...................))))))))))))))).)))(((........))).......................... (-17.78 = -18.64 + 0.86)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:25:16 2006