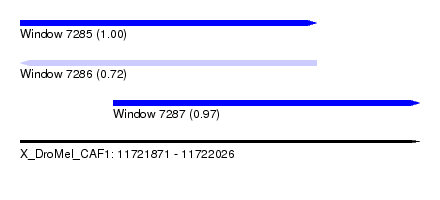
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,721,871 – 11,722,026 |
| Length | 155 |
| Max. P | 0.997010 |

| Location | 11,721,871 – 11,721,986 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 59.57 |
| Mean single sequence MFE | -35.02 |
| Consensus MFE | -16.22 |
| Energy contribution | -14.42 |
| Covariance contribution | -1.80 |
| Combinations/Pair | 1.71 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.46 |
| SVM decision value | 2.78 |
| SVM RNA-class probability | 0.997010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11721871 115 + 22224390 GAAUGACCCAGAUAUCACGGUUUAUCCCGC----CCAUGUGGCGCUUAUGACGCAGUUGUCUUAAACUCGAACU-CGAGCGGGCAAUUGCUGAUUACGAUUAACCACUGAUUCCUGGGUC ....(((((((..((((.((((.(((.(((----(.....))))........((((((((((....((((....-)))).)))))))))).......))).))))..))))..))))))) ( -44.10) >DroVir_CAF1 14738 96 + 1 AAAUACUCCAUAUCG-GUAUUAGAUCUCGCAAGAUUU-UUUGCGCAUAUGACGCAGCAUUCAUAAACUCAACGC-UGAGUGGAAUGUUGCUGAAU-----GGAU---------------- ......(((((.(((-(((..((((((....))))))-..))).........(((((((((....(((((....-))))).))))))))))))))-----))).---------------- ( -30.80) >DroSim_CAF1 14113 115 + 1 AAAUGGCCCAGAUAUCACGGUUUAUCCCGC----CCACGUGGCACUUAUGACGCAGUUGUCUUAAACUCGGACU-CGAGCGGGCAAUUGCUGAAUACGAUUGAUCACUGAUUCCUUGGCC ....((((.((..((((.((((.(((..((----(.....))).........((((((((((....((((....-)))).)))))))))).......))).))))..))))..)).)))) ( -35.60) >DroMoj_CAF1 30785 95 + 1 GAAUGGUCCAUAUCU-GUGUUAGAUCUCGCAAU-UCA-UUUGCGCUUAUGACGCAAUAUUCAUAAACUCAUCAUUUGAGUGGAAUAUUGCUGA-U-----ACAU---------------- ..............(-((((((.....(((((.-...-.)))))........(((((((((....(((((.....))))).))))))))))))-)-----))).---------------- ( -27.00) >DroAna_CAF1 21989 113 + 1 UAUUGACCCGGACAUCGCGUUCCAUUUCGU----CCCUGUGGCGUUUAUGACGCAGUGUUCCUAAACUCUAACC-UGAGCGGAGUACUGCCGA--ACAAUGGAUCCUUGAUUUCUGGGUU ....((((((((.((((...((((((.(((----(.....))))........((((((((((....(((.....-.))).))))))))))...--..))))))....)))).)))))))) ( -37.60) >consensus AAAUGACCCAGAUAUCGCGUUAGAUCUCGC____CCA_GUGGCGCUUAUGACGCAGUAUUCAUAAACUCAAACU_UGAGCGGAAUAUUGCUGA_UAC_AUGGAUC__UGAUU_CU_GG__ ....................................................((((((((((....((((.....)))).)))))))))).............................. (-16.22 = -14.42 + -1.80)



| Location | 11,721,871 – 11,721,986 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 59.57 |
| Mean single sequence MFE | -32.08 |
| Consensus MFE | -11.34 |
| Energy contribution | -10.38 |
| Covariance contribution | -0.96 |
| Combinations/Pair | 1.62 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.35 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11721871 115 - 22224390 GACCCAGGAAUCAGUGGUUAAUCGUAAUCAGCAAUUGCCCGCUCG-AGUUCGAGUUUAAGACAACUGCGUCAUAAGCGCCACAUGG----GCGGGAUAAACCGUGAUAUCUGGGUCAUUC ((((((((.((((.(((((.((((((((.....)))))(((((((-.((....(((((.(((......))).)))))...)).)))----))))))).))))))))).)))))))).... ( -47.40) >DroVir_CAF1 14738 96 - 1 ----------------AUCC-----AUUCAGCAACAUUCCACUCA-GCGUUGAGUUUAUGAAUGCUGCGUCAUAUGCGCAAA-AAAUCUUGCGAGAUCUAAUAC-CGAUAUGGAGUAUUU ----------------.(((-----((((.(((.(((((.((((.-.....))))....))))).)))........(((((.-.....)))))...........-.)).)))))...... ( -23.50) >DroSim_CAF1 14113 115 - 1 GGCCAAGGAAUCAGUGAUCAAUCGUAUUCAGCAAUUGCCCGCUCG-AGUCCGAGUUUAAGACAACUGCGUCAUAAGUGCCACGUGG----GCGGGAUAAACCGUGAUAUCUGGGCCAUUU ((((.(((.((((.....((((.((.....)).))))((((((((-.((..(..((((.(((......))).))))..).)).)))----)))))........)))).))).)))).... ( -33.70) >DroMoj_CAF1 30785 95 - 1 ----------------AUGU-----A-UCAGCAAUAUUCCACUCAAAUGAUGAGUUUAUGAAUAUUGCGUCAUAAGCGCAAA-UGA-AUUGCGAGAUCUAACAC-AGAUAUGGACCAUUC ----------------..((-----(-((.(((((((((.(((((.....)))))....)))))))))((......(((((.-...-.))))).......))..-.)))))......... ( -23.32) >DroAna_CAF1 21989 113 - 1 AACCCAGAAAUCAAGGAUCCAUUGU--UCGGCAGUACUCCGCUCA-GGUUAGAGUUUAGGAACACUGCGUCAUAAACGCCACAGGG----ACGAAAUGGAACGCGAUGUCCGGGUCAAUA .((((.((.(((..(..((((((..--((((((((..(((((((.-.....))))...)))..)))))..........((....))----.))))))))).)..))).)).))))..... ( -32.50) >consensus __CC_AG_AAUCA__GAUCAAU_GUA_UCAGCAAUAGUCCGCUCA_AGUUCGAGUUUAAGAAAACUGCGUCAUAAGCGCCAC_UGG____GCGAGAUAAAACGCGAUAUCUGGGCCAUUC ....................................(((((((((.....)))))...........((((.....))))................................))))..... (-11.34 = -10.38 + -0.96)

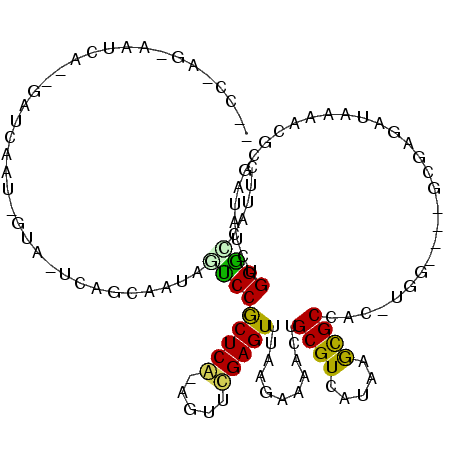

| Location | 11,721,907 – 11,722,026 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 73.95 |
| Mean single sequence MFE | -34.80 |
| Consensus MFE | -30.18 |
| Energy contribution | -29.87 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969881 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11721907 119 + 22224390 GGCGCUUAUGACGCAGUUGUCUUAAACUCGAACUCGAGCGGGCAAUUGCUGAUUACGAUUAACCACUGAUUCCUGGGUCGCUGCUUCGUGGCCGUCGUCGGUUCCAUUUAUCAACUAUU ((.(((.(((((((((((((((....((((....)))).))))))))))..........................((((((......))))))))))).))).)).............. ( -38.00) >DroSim_CAF1 14149 119 + 1 GGCACUUAUGACGCAGUUGUCUUAAACUCGGACUCGAGCGGGCAAUUGCUGAAUACGAUUGAUCACUGAUUCCUUGGCCGCUGCUUCGUGGCCGUCGUCGGUUCCAUUUAUCAACUAUU ((.(((.(((((((((((((((....((((....)))).))))))))))..........................((((((......))))))))))).))).)).............. ( -39.50) >DroAna_CAF1 22025 108 + 1 GGCGUUUAUGACGCAGUGUUCCUAAACUCUAACCUGAGCGGAGUACUGCCGA--ACAAUGGAUCCUUGAUUUCUGGGUUGCAGUUGCUAGGUCGUU-----UACCUUUUAACCAC---- ((.((..(((((((((((((((....(((......))).))))))))))...--.((((.(((((.........)))))...))))....))))).-----.)))).........---- ( -26.90) >consensus GGCGCUUAUGACGCAGUUGUCUUAAACUCGAACUCGAGCGGGCAAUUGCUGA_UACGAUUGAUCACUGAUUCCUGGGUCGCUGCUUCGUGGCCGUCGUCGGUUCCAUUUAUCAACUAUU ((.(((.(((((((((((((((....((((....)))).))))))))))..........................((((((......))))))))))).))).)).............. (-30.18 = -29.87 + -0.31)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:58 2006