
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,657,370 – 11,657,503 |
| Length | 133 |
| Max. P | 0.832793 |

| Location | 11,657,370 – 11,657,485 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 91.92 |
| Mean single sequence MFE | -49.18 |
| Consensus MFE | -43.42 |
| Energy contribution | -44.67 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.88 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.522542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
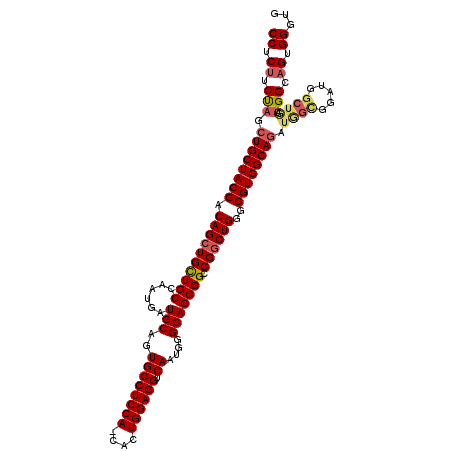
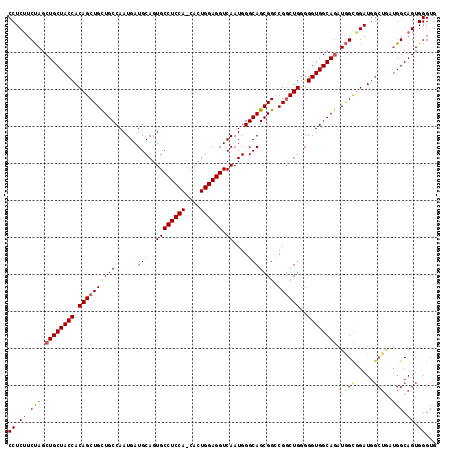
>X_DroMel_CAF1 11657370 115 - 22224390 CCUCUUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA-CACUGGAGGUCAAUGGGCAGCGGCCGGCUGGGGGUGGCAGAUGGCGGAUGGCUGAUGGCAGUGGGUG (((((.(((.((((((((.((((((((((......(((..(((((((.-....))))).))....))))))).))))))..)))))))).))).)))..((((....))))))... ( -51.30) >DroSec_CAF1 30567 116 - 1 CCUCUUCCAGCUGCUACCACAGCUGUUGCCAAUGAUGCAGUGCCUCCACCACUGGAGGUCAAUGGGCAGCGGCCGGCUGGGGGUGGCAAAUAGUGGAUGGCUGAUGGCAGUGGGUG ((.((.(((...((((((((.(((((((((..((.......(((((((....)))))))))...)))))))))..(((......))).....)))).))))...))).)).))... ( -47.91) >DroEre_CAF1 34682 108 - 1 CCUCGUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA-CACUGGAGGUCAAUGGGCAGCGGGCGACUGGGGGUGGCAGAUGGCUGAU-------GGCAGUGGGUG ((.(((((.((..(..(((..(((((((((..((.......((((((.-....))))))))...)))))).)))...)))..)..)))))))((((..-------..))))))... ( -46.41) >DroYak_CAF1 29544 115 - 1 CCUCUUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA-CACUGGAGGUCAAUGGGCAGCGGACGGCUGGGGGUGGCAGAUGGCUGAUGGCUAAUGGCAGUGGGUG ((.((.(((.((((((((.((((((((((......(((..(((((((.-....))))).))....))))))).))))))..)))))))).((((.....)))).))).)).))... ( -51.10) >consensus CCUCUUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA_CACUGGAGGUCAAUGGGCAGCGGCCGGCUGGGGGUGGCAGAUGGCGGAUGGCUGAUGGCAGUGGGUG ((.((.(((.((((((((.((((((((((......(((..((((((((....)))))).))....))))))).))))))..)))))))).((((.....)))).))).)).))... (-43.42 = -44.67 + 1.25)

| Location | 11,657,409 – 11,657,503 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 95.06 |
| Mean single sequence MFE | -40.93 |
| Consensus MFE | -37.52 |
| Energy contribution | -37.34 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802866 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
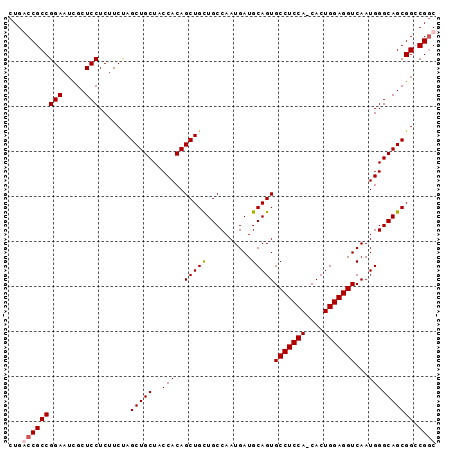
>X_DroMel_CAF1 11657409 94 + 22224390 GCCGGCCGCUGCCCAUUGACCUCCAGUG-UGGAGGCACUGCAUCAUUGGCAGCAGCUGUGGUAGCAGCUAGAAGAGGAGCGAUUCCGGCGGCCAG ((((.((((((((((.((.(((((....-.))))))).)).......))))))((((((....))))))......((((...))))))))))... ( -40.81) >DroSec_CAF1 30606 95 + 1 GCCGGCCGCUGCCCAUUGACCUCCAGUGGUGGAGGCACUGCAUCAUUGGCAACAGCUGUGGUAGCAGCUGGAAGAGGAGCAAUUCCGGCGGUCAG ...((((((((...((((.((((((((((((.((...)).))))))))....(((((((....)))))))...))))..))))..)))))))).. ( -43.80) >DroEre_CAF1 34714 94 + 1 GUCGCCCGCUGCCCAUUGACCUCCAGUG-UGGAGGCACUGCAUCAUUGGCAGCAGCUGUGGUAGCAGCUAGACGAGGAGCAAUUCCGGCGGUCAG ((((((.((((((((.((.(((((....-.))))))).)).......))))))((((((....))))))......((((...))))))))).).. ( -38.11) >DroYak_CAF1 29583 94 + 1 GCCGUCCGCUGCCCAUUGACCUCCAGUG-UGGAGGCACUGCAUCAUUGGCAGCAGCUGUGGUAGCAGCUAGAAGAGGAGCGAUUCCGGCGGCCAG ((((((.((((((((.((.(((((....-.))))))).)).......))))))((((((....))))))......((((...))))))))))... ( -41.01) >consensus GCCGGCCGCUGCCCAUUGACCUCCAGUG_UGGAGGCACUGCAUCAUUGGCAGCAGCUGUGGUAGCAGCUAGAAGAGGAGCAAUUCCGGCGGCCAG ((((.((((((((((.((.((((((....)))))))).)).......))))))((((((....))))))......((((...))))))))))... (-37.52 = -37.34 + -0.19)



| Location | 11,657,409 – 11,657,503 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 95.06 |
| Mean single sequence MFE | -37.85 |
| Consensus MFE | -34.89 |
| Energy contribution | -35.45 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.832793 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11657409 94 - 22224390 CUGGCCGCCGGAAUCGCUCCUCUUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA-CACUGGAGGUCAAUGGGCAGCGGCCGGC ((((((...(((.....))).......(((((..(((..(((((..........)))))((((((.-....))))))...)))))))))))))). ( -40.30) >DroSec_CAF1 30606 95 - 1 CUGACCGCCGGAAUUGCUCCUCUUCCAGCUGCUACCACAGCUGUUGCCAAUGAUGCAGUGCCUCCACCACUGGAGGUCAAUGGGCAGCGGCCGGC ......(((((..((((((......((((((......))))))((((.......)))).(((((((....)))))))....))))))...))))) ( -36.10) >DroEre_CAF1 34714 94 - 1 CUGACCGCCGGAAUUGCUCCUCGUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA-CACUGGAGGUCAAUGGGCAGCGGGCGAC .....(((((((.....))).......(((((..(((..(((((..........)))))((((((.-....))))))...)))))))).)))).. ( -36.60) >DroYak_CAF1 29583 94 - 1 CUGGCCGCCGGAAUCGCUCCUCUUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA-CACUGGAGGUCAAUGGGCAGCGGACGGC ...(((((((((.....))).......(((((..(((..(((((..........)))))((((((.-....))))))...)))))))))).)))) ( -38.40) >consensus CUGACCGCCGGAAUCGCUCCUCUUCUAGCUGCUACCACAGCUGCUGCCAAUGAUGCAGUGCCUCCA_CACUGGAGGUCAAUGGGCAGCGGCCGGC ...(((((((((.....))).......(((((..(((..(((((..........)))))(((((((....)))))))...)))))))))).)))) (-34.89 = -35.45 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:41 2006