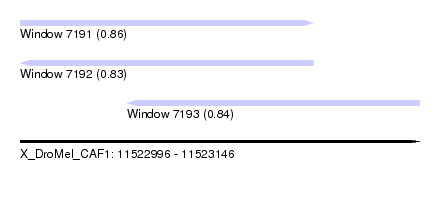
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,522,996 – 11,523,146 |
| Length | 150 |
| Max. P | 0.861923 |

| Location | 11,522,996 – 11,523,106 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.57 |
| Mean single sequence MFE | -37.37 |
| Consensus MFE | -18.98 |
| Energy contribution | -21.85 |
| Covariance contribution | 2.87 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11522996 110 + 22224390 ACAGUCGGUUAAUCACAACAAUACGGCGAUCUGGCAGAUCAGUUGGGGGGAUCUGCCACU------GCCACUGCCAUUGCUGGUGC----UAUUGCUUUAGGAAGUGGCAAACCCCUAAU ...((((................))))((((.....)))).((((((((....(((((((------.((...((((....))))((----....))....)).)))))))..)))))))) ( -38.89) >DroSec_CAF1 4387 110 + 1 ACAGUCGGUUAAUCACAACAAUACGGCGAUCUGGCAGAUCAGUUGGGGGGAUCUGCCACU------GCCACUGCCAUUGCUGGUGC----UAUUGCUUUAGGAAGUGGCAAACCCCUAAU ...((((................))))((((.....)))).((((((((....(((((((------.((...((((....))))((----....))....)).)))))))..)))))))) ( -38.89) >DroSim_CAF1 5926 98 + 1 ACAGUCGGUUAAUCACAACAAUACGGCGAUCUGGCAGAUCAGUUGGGGGGAUCUG------------------CCAUUGCUGGUGC----UAUUGCUUUAGGAAGUGGCAAACCCCUAAU ...((((................))))((((.....)))).((((((((....((------------------(((((.((((.((----....)).))))..)))))))..)))))))) ( -33.79) >DroYak_CAF1 4712 108 + 1 --------UUAAUU--AACAAUUCAGCGAUUUGGCAGAUCAACUAU--GCAUCUGCCACUGUCAUUGCCACUGCCAUUGGUGGUGCGCUACACUGCUGUAGGAAGUGGCAAACCCCUAAU --------......--...........(((.((((((((((....)--).))))))))..))).((((((((.((..(((..(((.....)))..)))..)).))))))))......... ( -37.90) >consensus ACAGUCGGUUAAUCACAACAAUACGGCGAUCUGGCAGAUCAGUUGGGGGGAUCUGCCACU______GCCACUGCCAUUGCUGGUGC____UAUUGCUUUAGGAAGUGGCAAACCCCUAAU ......((((.....................(((((((((.........)))))))))........((((((.(((..((.((((.....))))))..).)).)))))).))))...... (-18.98 = -21.85 + 2.87)



| Location | 11,522,996 – 11,523,106 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.57 |
| Mean single sequence MFE | -33.84 |
| Consensus MFE | -22.29 |
| Energy contribution | -23.60 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.825057 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11522996 110 - 22224390 AUUAGGGGUUUGCCACUUCCUAAAGCAAUA----GCACCAGCAAUGGCAGUGGC------AGUGGCAGAUCCCCCCAACUGAUCUGCCAGAUCGCCGUAUUGUUGUGAUUAACCGACUGU ..(((.(((((((((((.......((....----)).(((....))).))))))------).(((((((((.........)))))))))((((((.........))))))))))..))). ( -36.70) >DroSec_CAF1 4387 110 - 1 AUUAGGGGUUUGCCACUUCCUAAAGCAAUA----GCACCAGCAAUGGCAGUGGC------AGUGGCAGAUCCCCCCAACUGAUCUGCCAGAUCGCCGUAUUGUUGUGAUUAACCGACUGU ..(((.(((((((((((.......((....----)).(((....))).))))))------).(((((((((.........)))))))))((((((.........))))))))))..))). ( -36.70) >DroSim_CAF1 5926 98 - 1 AUUAGGGGUUUGCCACUUCCUAAAGCAAUA----GCACCAGCAAUGG------------------CAGAUCCCCCCAACUGAUCUGCCAGAUCGCCGUAUUGUUGUGAUUAACCGACUGU .(((((((........))))))).((((((----(.((..((..(((------------------((((((.........)))))))))....)).)).))))))).............. ( -26.30) >DroYak_CAF1 4712 108 - 1 AUUAGGGGUUUGCCACUUCCUACAGCAGUGUAGCGCACCACCAAUGGCAGUGGCAAUGACAGUGGCAGAUGC--AUAGUUGAUCUGCCAAAUCGCUGAAUUGUU--AAUUAA-------- .((((.(((((((((((.((.......(((.....))).......)).))))))).......((((((((.(--(....)))))))))))))).))))......--......-------- ( -35.64) >consensus AUUAGGGGUUUGCCACUUCCUAAAGCAAUA____GCACCAGCAAUGGCAGUGGC______AGUGGCAGAUCCCCCCAACUGAUCUGCCAGAUCGCCGUAUUGUUGUGAUUAACCGACUGU .(((((((........)))))))(((((((....((.(((....)))...............(((((((((.........)))))))))....))..)))))))................ (-22.29 = -23.60 + 1.31)


| Location | 11,523,036 – 11,523,146 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.37 |
| Mean single sequence MFE | -32.11 |
| Consensus MFE | -21.16 |
| Energy contribution | -23.46 |
| Covariance contribution | 2.30 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.842465 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11523036 110 - 22224390 CAUCGAACUAUUCGCAGCUGUGAUGCAAGCUUCGAUCUCUAUUAGGGGUUUGCCACUUCCUAAAGCAAUA----GCACCAGCAAUGGCAGUGGC------AGUGGCAGAUCCCCCCAACU .((((((((..((((....))))....)).))))))........(((((((((((((.((....((....----)).(((....)))....)).------)))))))))))))....... ( -39.30) >DroSec_CAF1 4427 110 - 1 CAUCGAACUAUUCGCAGCUGUGAUGCAAGCCUCGAUCUCUAUUAGGGGUUUGCCACUUCCUAAAGCAAUA----GCACCAGCAAUGGCAGUGGC------AGUGGCAGAUCCCCCCAACU .(((((.((..((((....))))....))..)))))........(((((((((((((.((....((....----)).(((....)))....)).------)))))))))))))....... ( -37.50) >DroSim_CAF1 5966 98 - 1 CAUCGAACUAUUCGCAGCUGUGAUGCAAGCCUCGAUCUCUAUUAGGGGUUUGCCACUUCCUAAAGCAAUA----GCACCAGCAAUGG------------------CAGAUCCCCCCAACU .(((((.((..((((....))))....))..)))))........(((((((((((.((.((...((....----))...)).)))))------------------))))))))....... ( -25.70) >DroEre_CAF1 4406 85 - 1 CAUCGAACUAUUCGCAGCCGUGAUGCAAGUCUCGAUCUCUAUUAGGGGUUUGCCACUUCCUAAAGCAAAG----GCAC---------------------CAGUGGCAGAU---------- .((((((((..((((....))))....))).)))))(((.....)))((((((((((.(((.......))----)...---------------------.))))))))))---------- ( -23.60) >DroYak_CAF1 4742 118 - 1 CAUCGAACUAUUCGCAGCUGUGAUGCAAGCUUCGAUCUCUAUUAGGGGUUUGCCACUUCCUACAGCAGUGUAGCGCACCACCAAUGGCAGUGGCAAUGACAGUGGCAGAUGC--AUAGUU .....((((((..(((.((((.((.((......((((((.....))))))(((((((.((.......(((.....))).......)).))))))).))...)).)))).)))--)))))) ( -34.44) >consensus CAUCGAACUAUUCGCAGCUGUGAUGCAAGCCUCGAUCUCUAUUAGGGGUUUGCCACUUCCUAAAGCAAUA____GCACCAGCAAUGGCAGUGGC______AGUGGCAGAUCCCCCCAACU .(((((.....((((....))))........)))))........(((((((((((((.......((........)).(((....))).............)))))))))))))....... (-21.16 = -23.46 + 2.30)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:23:33 2006