
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,368,766 – 11,368,857 |
| Length | 91 |
| Max. P | 0.998425 |

| Location | 11,368,766 – 11,368,857 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 75.16 |
| Mean single sequence MFE | -27.48 |
| Consensus MFE | -21.24 |
| Energy contribution | -20.80 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982649 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11368766 91 + 22224390 ----------------UGCUUUGUGGGUCAGCUCAUUGAAAAGUUGACUGCCAAUUGCAGUUAACCAAUUAGCAAUACUCGACUCUU-UGUUGCUGUUGUUGUUUUGU------- ----------------...((.(((((....))))).))...(((((((((.....)))))))))(((.((((((((..((((....-.)))).))))))))..))).------- ( -24.10) >DroSim_CAF1 10729 85 + 1 ----------------UGCUUUGUGGGUCAGCUCAUUGAAAAGUUGACUGCCAAUUGCAGUUAACCAAUUAGCAAUACUCGACUCUU-UGUUGCUGUUGUUG------------- ----------------...((.(((((....))))).))...(((((((((.....)))))))))....((((((((..((((....-.)))).))))))))------------- ( -22.60) >DroYak_CAF1 34727 98 + 1 ----------------UGCUUUGUGGGUCAGCUCAUUGAAAAGUUGACUGCCAAUCGCAGUUAACCAAUUAGCAAUACUCGGCUCUU-UGUUGCUGUUGUUAUUUUGUUGUUGUU ----------------...((.(((((....))))).))...(((((((((.....)))))))))((((.(((((.((.((((....-....))))..))....))))))))).. ( -24.80) >DroAna_CAF1 21096 101 + 1 UUGUUUGCGGUGUAAGCGCUUUGUGGGUCAGCUCAUUGAAAAGCUGACUGCCAAUUGCAGCUAACCAAUUAGCGAUGGCUGACUGUUGCGCCGCUGCUGGC-------------- ..((..(((((((((((((.(((..((((((((........))))))))..)))..)).)))...((.(((((....))))).)).)))))))).))....-------------- ( -38.40) >consensus ________________UGCUUUGUGGGUCAGCUCAUUGAAAAGUUGACUGCCAAUUGCAGUUAACCAAUUAGCAAUACUCGACUCUU_UGUUGCUGUUGUUG_____________ .................((.(((..((((((((........))))))))..)))..)).......((((.(((((((...........)))))))))))................ (-21.24 = -20.80 + -0.44)


| Location | 11,368,766 – 11,368,857 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 75.16 |
| Mean single sequence MFE | -23.62 |
| Consensus MFE | -17.46 |
| Energy contribution | -17.77 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.74 |
| SVM decision value | 3.10 |
| SVM RNA-class probability | 0.998425 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
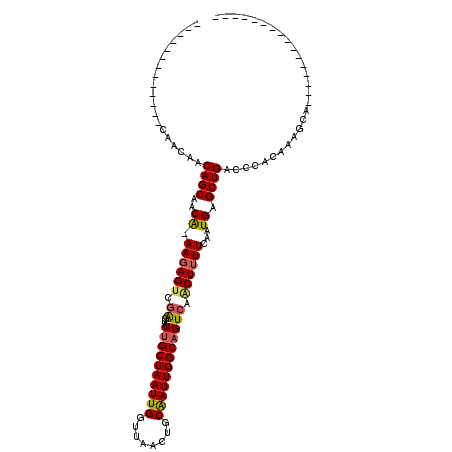
>X_DroMel_CAF1 11368766 91 - 22224390 -------ACAAAACAACAACAGCAACA-AAGAGUCGAGUAUUGCUAAUUGGUUAACUGCAAUUGGCAGUCAACUUUUCAAUGAGCUGACCCACAAAGCA---------------- -------............((((..((-..(((..(((((((((((((((........)))))))))))..)))))))..)).))))............---------------- ( -19.20) >DroSim_CAF1 10729 85 - 1 -------------CAACAACAGCAACA-AAGAGUCGAGUAUUGCUAAUUGGUUAACUGCAAUUGGCAGUCAACUUUUCAAUGAGCUGACCCACAAAGCA---------------- -------------......((((..((-..(((..(((((((((((((((........)))))))))))..)))))))..)).))))............---------------- ( -19.20) >DroYak_CAF1 34727 98 - 1 AACAACAACAAAAUAACAACAGCAACA-AAGAGCCGAGUAUUGCUAAUUGGUUAACUGCGAUUGGCAGUCAACUUUUCAAUGAGCUGACCCACAAAGCA---------------- ...................((((..((-..(((..(((((((((((((((........)))))))))))..)))))))..)).))))............---------------- ( -18.90) >DroAna_CAF1 21096 101 - 1 --------------GCCAGCAGCGGCGCAACAGUCAGCCAUCGCUAAUUGGUUAGCUGCAAUUGGCAGUCAGCUUUUCAAUGAGCUGACCCACAAAGCGCUUACACCGCAAACAA --------------(((((..(((((....((((.(((....))).))))....)))))..))))).((((((((......)))))))).......(((.......)))...... ( -37.20) >consensus _____________CAACAACAGCAACA_AAGAGUCGAGUAUUGCUAAUUGGUUAACUGCAAUUGGCAGUCAACUUUUCAAUGAGCUGACCCACAAAGCA________________ ...................((((..((.((((((.((...((((((((((........)))))))))))).))))))...)).))))............................ (-17.46 = -17.77 + 0.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:21:54 2006