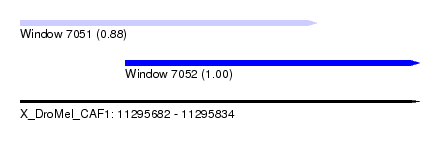
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,295,682 – 11,295,834 |
| Length | 152 |
| Max. P | 0.998929 |

| Location | 11,295,682 – 11,295,795 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 84.39 |
| Mean single sequence MFE | -27.28 |
| Consensus MFE | -21.22 |
| Energy contribution | -21.97 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.877172 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
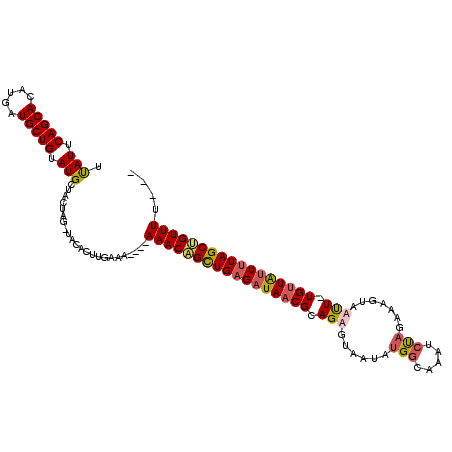
>X_DroMel_CAF1 11295682 113 + 22224390 UCAUUCAGCACAUGAUGCUGUAUGCUACUAGUUACACUUGAAAAAAAAAACAGCUGGGAUAACGCAGAGUAAUAUGGCAAAUCUAGAAAGUAUUU-UGUUAUUGUAGCUGUUUU--- .(((.(((((.....))))).)))......................((((((((((.((((..((((((((...(((.....))).....)))))-))))))).))))))))))--- ( -27.80) >DroSim_CAF1 20986 113 + 1 UUAUUCAGCACAUGAUGCUGUAUGCUACUAG-UACACUUGAAAA--AAAACAGCUGAGAUAACGCAGAGUAAUAUGGCAAAUCUAGAAUGUAUCU-UGUUAUUUUAGCUGUUUUUUG .(((.(((((.....))))).))).......-..........((--((((((((((((((((((.(((......(((.....))).......)))-)))))))))))))))))))). ( -30.92) >DroEre_CAF1 25500 108 + 1 UUAUUCAGCACAUGAUGCUGUAUGCUACUAG-UACACUUGAAA----AAACAGCUGAGAUAACGCAGAGCAAAAAGGCAAUUGCAUAAAGUAAUU-UGUUGCUUUAGCAGUUUU--- ...((((((...((.(((((........)))-)))).((....----))...))))))..........((......))((((((.((((((((..-..))))))))))))))..--- ( -21.70) >DroYak_CAF1 26325 109 + 1 UUAUUCAGCACAUGAUGCUGUAUGCUGCCAG-UACACUUGGAG----AAACGGUUGAGAUAACGCAGAGUAAUAUGGCAAUGCUAGAAAGUAAUUUUGGUGUUUUAGCCGUUUU--- .(((.(((((.....))))).)))...((((-.....)))).(----((((((((((((((.(.((((((.((.((((...))))....)).))))))))))))))))))))))--- ( -28.70) >consensus UUAUUCAGCACAUGAUGCUGUAUGCUACUAG_UACACUUGAAA____AAACAGCUGAGAUAACGCAGAGUAAUAUGGCAAAUCUAGAAAGUAAUU_UGUUAUUUUAGCUGUUUU___ .(((.(((((.....))))).))).......................(((((((((((((((((.(((......(((.....))).......))).))))))))))))))))).... (-21.22 = -21.97 + 0.75)

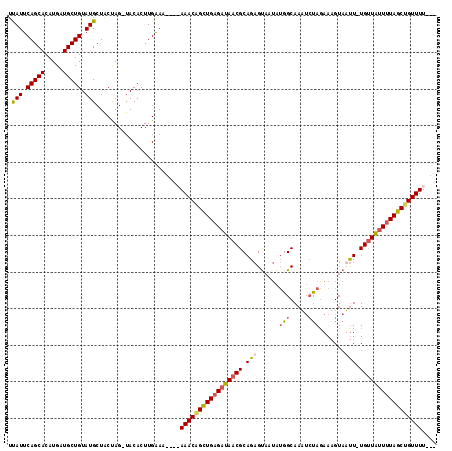
| Location | 11,295,722 – 11,295,834 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 79.38 |
| Mean single sequence MFE | -28.36 |
| Consensus MFE | -18.78 |
| Energy contribution | -19.97 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.20 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.66 |
| SVM decision value | 3.29 |
| SVM RNA-class probability | 0.998929 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
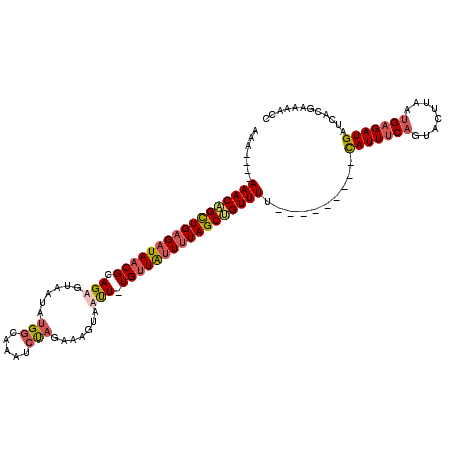
>X_DroMel_CAF1 11295722 112 + 22224390 AAAAAAAAAACAGCUGGGAUAACGCAGAGUAAUAUGGCAAAUCUAGAAAGUAUUU-UGUUAUUGUAGCUGUUUU----UUUUUCAUUUCAGUACUUAAUGAGAUGAUCACGAAAAGC ..((((((((((((((.((((..((((((((...(((.....))).....)))))-))))))).))))))))))----))))((((((((........))))))))........... ( -32.50) >DroSim_CAF1 21025 114 + 1 AAAA--AAAACAGCUGAGAUAACGCAGAGUAAUAUGGCAAAUCUAGAAUGUAUCU-UGUUAUUUUAGCUGUUUUUUGUUUUUUCAUUUCAGUACUUAAUGAGAUGAUCACGAAAAGC ..((--((((((((((((((((((.(((......(((.....))).......)))-))))))))))))))))))))((((((((((((((........)))))))).....)))))) ( -33.82) >DroEre_CAF1 25539 103 + 1 AAA----AAACAGCUGAGAUAACGCAGAGCAAAAAGGCAAUUGCAUAAAGUAAUU-UGUUGCUUUAGCAGUUUU---------UAUUUCAGUACUUAAUGCGAUGAUCACGAAACCC ...----.....((((((((((......((......))((((((.((((((((..-..)))))))))))))).)---------)))))))))......................... ( -22.90) >DroYak_CAF1 26364 104 + 1 GAG----AAACGGUUGAGAUAACGCAGAGUAAUAUGGCAAUGCUAGAAAGUAAUUUUGGUGUUUUAGCCGUUUU---------UAUUUCAGUACUUAACGAGAUGAUCACGAAUUCC .((----((((((((((((((.(.((((((.((.((((...))))....)).))))))))))))))))))))))---------)(((((.((.....)))))))............. ( -24.20) >consensus AAA____AAACAGCUGAGAUAACGCAGAGUAAUAUGGCAAAUCUAGAAAGUAAUU_UGUUAUUUUAGCUGUUUU_________CAUUUCAGUACUUAAUGAGAUGAUCACGAAAACC .......(((((((((((((((((.(((......(((.....))).......))).)))))))))))))))))..........(((((((........)))))))............ (-18.78 = -19.97 + 1.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:21:20 2006