
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,236,434 – 11,236,550 |
| Length | 116 |
| Max. P | 0.699734 |

| Location | 11,236,434 – 11,236,550 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 78.04 |
| Mean single sequence MFE | -23.59 |
| Consensus MFE | -15.17 |
| Energy contribution | -15.54 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.64 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.529613 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
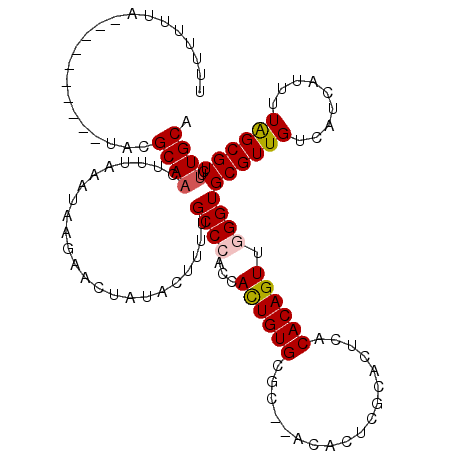
>X_DroMel_CAF1 11236434 116 + 22224390 UUUUUUUACGCAUAGAACUACGCAUAUAGAACUAAGUUCUAUAUUUUUGCCCACCACUGUGCGC--ACACUCGCACUCACACAGUUUGGUGCGUUGUCAUCAUUCUAGCGUCUUUGCA .........(((.(((....((((((((((((...)))))))))....(((((..((((((.((--......)).....)))))).))).))...............)))))).))). ( -28.60) >DroSec_CAF1 23349 101 + 1 UUUUUUUU---------UUACGCAACUUUGAAUAAGUUUUAUAUUUUUGCCCACCACUGUGCGC--ACAC------UCACACAGUUAGGUGCGUUGUCAACAUUUUAGCGUCUUUGCA ........---------..(((((((((.....)))))..........((.((((((((((.(.--....------.).))))))..)))).)).............))))....... ( -18.60) >DroEre_CAF1 23167 106 + 1 UUUUUUUC----------CCCGCAACUUUCAAUAGGAACUAUACUUUUGCCCUACACUGUGCGC--ACACUCGCACUCACACAGUUGGGUGCGUUGUCAUCAUUUUGGCGUCUUUGCA ........----------...((((.......((((..............))))......((((--(((..((((((((......)))))))).)))((......))))))..)))). ( -22.74) >DroYak_CAF1 15350 107 + 1 UCUUUUUA-----------GCGCAACUCUAAAUGGGAACUAUACUAUUGCCCUCCAUUGUGCACACACACUCGCACUCACACAGUUGGGUGCGUUGUCAUCAUUUUAGCGUCUUUGCA ........-----------((((((.....(((((........)))))........))))))....(((..((((((((......)))))))).)))..........(((....))). ( -24.42) >consensus UUUUUUUA__________UACGCAACUUUAAAUAAGAACUAUACUUUUGCCCACCACUGUGCGC__ACACUCGCACUCACACAGUUGGGUGCGUUGUCAUCAUUUUAGCGUCUUUGCA .....................((((.......................((((...((((((..................)))))).))))((((((.........))))))..)))). (-15.17 = -15.54 + 0.37)


| Location | 11,236,434 – 11,236,550 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 78.04 |
| Mean single sequence MFE | -28.08 |
| Consensus MFE | -16.56 |
| Energy contribution | -17.12 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.699734 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
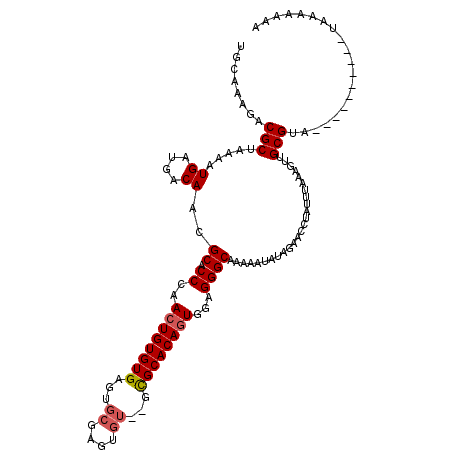
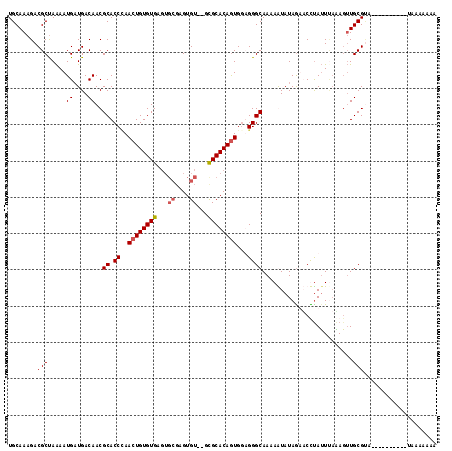
>X_DroMel_CAF1 11236434 116 - 22224390 UGCAAAGACGCUAGAAUGAUGACAACGCACCAAACUGUGUGAGUGCGAGUGU--GCGCACAGUGGUGGGCAAAAAUAUAGAACUUAGUUCUAUAUGCGUAGUUCUAUGCGUAAAAAAA .......((((((((((.(((.(...((.(((.((((((((...((....))--.))))))))..)))))....(((((((((...))))))))))))).)))))).))))....... ( -38.70) >DroSec_CAF1 23349 101 - 1 UGCAAAGACGCUAAAAUGUUGACAACGCACCUAACUGUGUGA------GUGU--GCGCACAGUGGUGGGCAAAAAUAUAAAACUUAUUCAAAGUUGCGUAA---------AAAAAAAA .......((((....(((((.....(.((((..((((((((.------....--.)))))))))))).)....)))))..(((((.....)))))))))..---------........ ( -25.60) >DroEre_CAF1 23167 106 - 1 UGCAAAGACGCCAAAAUGAUGACAACGCACCCAACUGUGUGAGUGCGAGUGU--GCGCACAGUGUAGGGCAAAAGUAUAGUUCCUAUUGAAAGUUGCGGG----------GAAAAAAA (((....((((..............(((((......))))).(((((.....--.))))).))))...))).........(((((..(....).....))----------)))..... ( -23.30) >DroYak_CAF1 15350 107 - 1 UGCAAAGACGCUAAAAUGAUGACAACGCACCCAACUGUGUGAGUGCGAGUGUGUGUGCACAAUGGAGGGCAAUAGUAUAGUUCCCAUUUAGAGUUGCGC-----------UAAAAAGA .((......))..............(((((......)))))(((((((((((....))))(((((.((((.........))))))))).....))))))-----------)....... ( -24.70) >consensus UGCAAAGACGCUAAAAUGAUGACAACGCACCCAACUGUGUGAGUGCGAGUGU__GCGCACAGUGGAGGGCAAAAAUAUAGAACCUAUUUAAAGUUGCGUA__________UAAAAAAA ........(((.....((....))..((.((..((((((((...((....))...))))))))...)))).........................))).................... (-16.56 = -17.12 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:20:54 2006