
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,141,475 – 11,141,589 |
| Length | 114 |
| Max. P | 0.955152 |

| Location | 11,141,475 – 11,141,589 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 91.70 |
| Mean single sequence MFE | -23.43 |
| Consensus MFE | -19.19 |
| Energy contribution | -19.25 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.821416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11141475 114 + 22224390 CA-UUUUAAGCAUCGUAACAUUUUCGACUGUAUAUUUUUGAUUGUUUUACCACUUUAUCUGCGCAUACAAAACCAAUUUGGUUCCCUUUGCCACUCGGGCGUAUGCUCGAUAAAA ..-...........((((.((..((((..........))))..)).))))...((((((.(.((((((..((((.....))))......(((.....))))))))).))))))). ( -23.00) >DroSec_CAF1 55944 114 + 1 UA-UUUGAAGCAUCGAAACAUUUUCUACUGUAUAUUUUUGAUUGUUUUACCACUUUAUCUGCGCAUACAAAACCAAUUUGGUUCCCUUUGCCACUCGGGCGUAUGCUCGAUAAAA ..-..(((((((((((((.((...(....)...)))))))).)))))))....((((((.(.((((((..((((.....))))......(((.....))))))))).))))))). ( -25.90) >DroEre_CAF1 56612 115 + 1 UUUUUUUAAAUGUCGAAACAUUUUCGACUGUAUAUUUUUGAUUGUUUUACCACUUUAUCUGCGCAUACAAAACCAAUUUGGUUGUCUUUGCCACUCGGGCGUAUACUCGAUAAAA ...........((((((.....)))))).........................((((((.(.(.((((..((((.....))))......(((.....))))))).).))))))). ( -20.00) >DroYak_CAF1 59639 111 + 1 ----UUUAAAUAACGAAAAAUUGUCGACUGCAUAUUUUUGAUUGUUUUACCACUUUAUCUGCGCAUACAAAACCAAUUUGGUUCCCUUUGCCACUCGGGCGUAUGCUCGAUAAAA ----...((((((((((((..(((.....)))..)))))).))))))......((((((.(.((((((..((((.....))))......(((.....))))))))).))))))). ( -24.80) >consensus UA_UUUUAAACAUCGAAACAUUUUCGACUGUAUAUUUUUGAUUGUUUUACCACUUUAUCUGCGCAUACAAAACCAAUUUGGUUCCCUUUGCCACUCGGGCGUAUGCUCGAUAAAA .......(((((((((((.................)))))).)))))......((((((.(.((((((..((((.....))))......(((.....))))))))).))))))). (-19.19 = -19.25 + 0.06)


| Location | 11,141,475 – 11,141,589 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 91.70 |
| Mean single sequence MFE | -25.62 |
| Consensus MFE | -24.24 |
| Energy contribution | -24.92 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955152 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
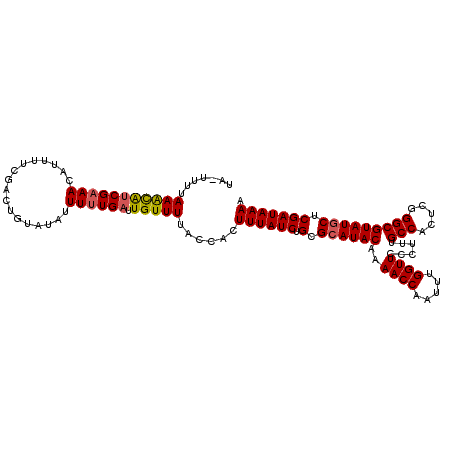
>X_DroMel_CAF1 11141475 114 - 22224390 UUUUAUCGAGCAUACGCCCGAGUGGCAAAGGGAACCAAAUUGGUUUUGUAUGCGCAGAUAAAGUGGUAAAACAAUCAAAAAUAUACAGUCGAAAAUGUUACGAUGCUUAAAA-UG (((((((..(((((((((.....)))....((((((.....))))))))))))...)))))))((((......))))..........((((.........))))........-.. ( -25.30) >DroSec_CAF1 55944 114 - 1 UUUUAUCGAGCAUACGCCCGAGUGGCAAAGGGAACCAAAUUGGUUUUGUAUGCGCAGAUAAAGUGGUAAAACAAUCAAAAAUAUACAGUAGAAAAUGUUUCGAUGCUUCAAA-UA .((((((..(((((((((.....)))....((((((.....))))))))))))...))))))(.(((((((((......................)))))...)))).)...-.. ( -24.35) >DroEre_CAF1 56612 115 - 1 UUUUAUCGAGUAUACGCCCGAGUGGCAAAGACAACCAAAUUGGUUUUGUAUGCGCAGAUAAAGUGGUAAAACAAUCAAAAAUAUACAGUCGAAAAUGUUUCGACAUUUAAAAAAA (((((((..(((((((((.....)))......((((.....))))..))))))...)))))))((((......))))..........((((((.....))))))........... ( -26.30) >DroYak_CAF1 59639 111 - 1 UUUUAUCGAGCAUACGCCCGAGUGGCAAAGGGAACCAAAUUGGUUUUGUAUGCGCAGAUAAAGUGGUAAAACAAUCAAAAAUAUGCAGUCGACAAUUUUUCGUUAUUUAAA---- .....(((((((((((((.....)))....((((((.....))))))))))))(((.......((((......))))......)))..))))...................---- ( -26.52) >consensus UUUUAUCGAGCAUACGCCCGAGUGGCAAAGGGAACCAAAUUGGUUUUGUAUGCGCAGAUAAAGUGGUAAAACAAUCAAAAAUAUACAGUCGAAAAUGUUUCGAUACUUAAAA_UA (((((((..(((((((((.....)))....((((((.....))))))))))))...)))))))((((......))))..........((((((.....))))))........... (-24.24 = -24.92 + 0.69)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:20:11 2006