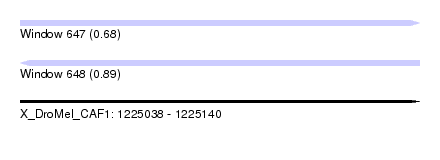
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,225,038 – 1,225,140 |
| Length | 102 |
| Max. P | 0.889220 |

| Location | 1,225,038 – 1,225,140 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 92.45 |
| Mean single sequence MFE | -33.76 |
| Consensus MFE | -29.93 |
| Energy contribution | -29.83 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.683635 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1225038 102 + 22224390 GCGUGGUGGCGUUGUGGCUGUUAUGCAAAUCGUGCGACUGGGAAAGCAGUUUACUUUUACUUUUCAAAUCGAUUAUAUGUGCACACAGC------CGCCAUUUCGUGC (((..(((((.....((((((..((((.......(((...((((((.((.......)).))))))...)))........)))).)))))------))))))..))).. ( -30.56) >DroSim_CAF1 2118 102 + 1 GCGUGGUGGCGGUUUGGCUGUUAUGCAAAUCGUGCGACUGGGAAAGCAGUUUACUUUUACUUUUCAAAUCGAUUAUAUGUGCACACAGC------CGCCAUUUCGUGC (((..(((((.....((((((..((((.......(((...((((((.((.......)).))))))...)))........)))).)))))------))))))..))).. ( -30.56) >DroYak_CAF1 1801 107 + 1 -UGGGGUGGCGUUGUGGCUGUUAUGCAAAUCGUGCGACUGGGAAAGCAGUUUACUUUUACUUUUCAAAUCGAUUAUAUGUGCACACAGUGCACAGCGCCAUUUCGUGC -.(..((((((((((((((((..((((.......(((...((((((.((.......)).))))))...)))........)))).))))).)))))))))))..).... ( -40.16) >consensus GCGUGGUGGCGUUGUGGCUGUUAUGCAAAUCGUGCGACUGGGAAAGCAGUUUACUUUUACUUUUCAAAUCGAUUAUAUGUGCACACAGC______CGCCAUUUCGUGC (((..((((((.....(((((..((((.......(((...((((((.((.......)).))))))...)))........)))).)))))......))))))..))).. (-29.93 = -29.83 + -0.11)

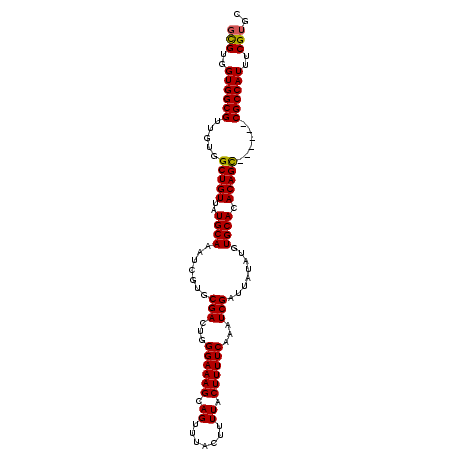
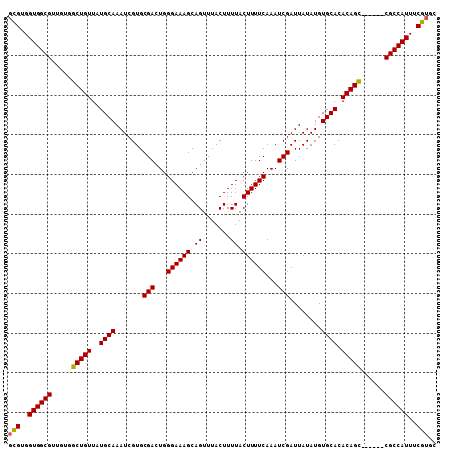
| Location | 1,225,038 – 1,225,140 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 92.45 |
| Mean single sequence MFE | -28.53 |
| Consensus MFE | -25.63 |
| Energy contribution | -25.97 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.889220 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1225038 102 - 22224390 GCACGAAAUGGCG------GCUGUGUGCACAUAUAAUCGAUUUGAAAAGUAAAAGUAAACUGCUUUCCCAGUCGCACGAUUUGCAUAACAGCCACAACGCCACCACGC ...((...(((((------((((((((((.((.....(((((.(.((((((.........)))))).).)))))....)).))))).)))))).....))))...)). ( -27.70) >DroSim_CAF1 2118 102 - 1 GCACGAAAUGGCG------GCUGUGUGCACAUAUAAUCGAUUUGAAAAGUAAAAGUAAACUGCUUUCCCAGUCGCACGAUUUGCAUAACAGCCAAACCGCCACCACGC ...((...(((((------((((((((((.((.....(((((.(.((((((.........)))))).).)))))....)).))))).)))))).....))))...)). ( -27.70) >DroYak_CAF1 1801 107 - 1 GCACGAAAUGGCGCUGUGCACUGUGUGCACAUAUAAUCGAUUUGAAAAGUAAAAGUAAACUGCUUUCCCAGUCGCACGAUUUGCAUAACAGCCACAACGCCACCCCA- ........(((((.((((..(((((((((.((.....(((((.(.((((((.........)))))).).)))))....)).))))).)))).)))).))))).....- ( -30.20) >consensus GCACGAAAUGGCG______GCUGUGUGCACAUAUAAUCGAUUUGAAAAGUAAAAGUAAACUGCUUUCCCAGUCGCACGAUUUGCAUAACAGCCACAACGCCACCACGC ........(((((......((((((((((.((.....(((((.(.((((((.........)))))).).)))))....)).))))).))))).....)))))...... (-25.63 = -25.97 + 0.33)
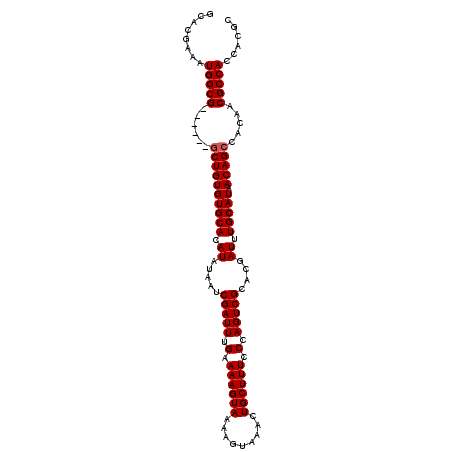


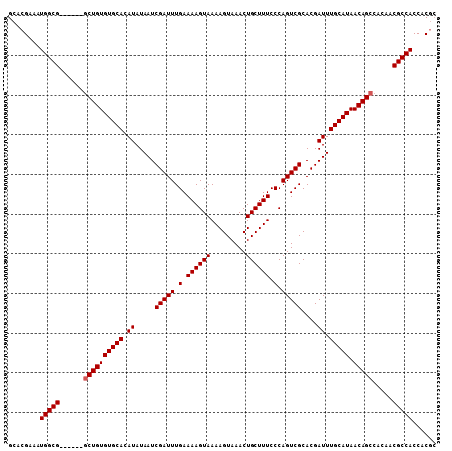
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:44:11 2006