
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,035,932 – 11,036,040 |
| Length | 108 |
| Max. P | 0.993505 |

| Location | 11,035,932 – 11,036,040 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 76.62 |
| Mean single sequence MFE | -35.73 |
| Consensus MFE | -19.69 |
| Energy contribution | -19.88 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792867 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11035932 108 + 22224390 GGAUUCCCCAUGUUGUUUCCCUUGGGGAAAUGGACCAAUUUU--CCCCAGGCAGUUGUGGUUCGCUGAGGUUGUCCCAGAAUGGGAGAUGGAAUGUGAAUCCUU--------UUUAUU ((((((..(((.(..(((((((((((((((.........)))--))))))...(((.(((..(((....).))..))).)))))))))..).))).))))))..--------...... ( -38.10) >DroSec_CAF1 23260 109 + 1 GAAUUACC-UCGUUGUUUCCCUUGGGGAAAUGGGCCCAUUUUUGCCCCUGGCAGUUGUGGAUCGCUGAGGUUGUCCCAAAAUGGGAGUUGGAAUGUGAAUCCUU--------UUUAUU .....(((-((...(((((((...))))))).(..((((..(((((...)))))..))))..)...)))))..((((.....))))...(((.......)))..--------...... ( -33.80) >DroSim_CAF1 15049 109 + 1 GGAUUCCC-UCGUUGUUUCCCUUGGGGAAAUGGGCCCAGUUUUGCCCCAGGCAGUUGUGGAUCGCUGAGGUUGUCCCAGAAUGGGAGUUGGAAUGUGAAUCCUU--------UUUAUU ((((((((-((...(((((((...))))))).(..(((...(((((...)))))...)))..)...))))...((((.....))))..........))))))..--------...... ( -35.60) >DroYak_CAF1 18381 102 + 1 --AU-------------UCCUCCAGGGAAAUGUGGUCAUUUC-CCUACGGGCAGAUGUGGGUCACUGAGGUUGCCCCAUAAUUGGAGAUGGAAUGAGAAUCCUUUUUCCUCUUUUAUU --..-------------..(((((((((((((....))))))-)))..((((((...(((....)))...)))))).......))))..((((.(((....))).))))......... ( -35.40) >consensus GGAUUCCC_UCGUUGUUUCCCUUGGGGAAAUGGGCCCAUUUU_GCCCCAGGCAGUUGUGGAUCGCUGAGGUUGUCCCAGAAUGGGAGAUGGAAUGUGAAUCCUU________UUUAUU .................(((((((((((((((....)))))...))))))....(((.((((.((....)).)))))))...))))...(((.......)))................ (-19.69 = -19.88 + 0.19)


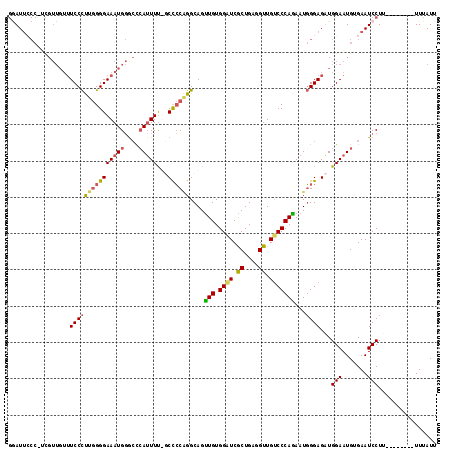
| Location | 11,035,932 – 11,036,040 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 76.62 |
| Mean single sequence MFE | -30.95 |
| Consensus MFE | -15.71 |
| Energy contribution | -16.90 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -3.09 |
| Structure conservation index | 0.51 |
| SVM decision value | 2.40 |
| SVM RNA-class probability | 0.993505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11035932 108 - 22224390 AAUAAA--------AAGGAUUCACAUUCCAUCUCCCAUUCUGGGACAACCUCAGCGAACCACAACUGCCUGGGG--AAAAUUGGUCCAUUUCCCCAAGGGAAACAACAUGGGGAAUCC ......--------..(((((...........((((.....))))...((((((((.........)).))))))--......)))))..(((((((..(....)....)))))))... ( -30.60) >DroSec_CAF1 23260 109 - 1 AAUAAA--------AAGGAUUCACAUUCCAACUCCCAUUUUGGGACAACCUCAGCGAUCCACAACUGCCAGGGGCAAAAAUGGGCCCAUUUCCCCAAGGGAAACAACGA-GGUAAUUC ......--------..(((.......)))...((((.....))))..(((((.(.(..(((....((((...))))....)))..)).((((((...))))))....))-)))..... ( -29.20) >DroSim_CAF1 15049 109 - 1 AAUAAA--------AAGGAUUCACAUUCCAACUCCCAUUCUGGGACAACCUCAGCGAUCCACAACUGCCUGGGGCAAAACUGGGCCCAUUUCCCCAAGGGAAACAACGA-GGGAAUCC ......--------..((((((..........((((.....))))...((((..............(((..(.......)..)))...((((((...))))))....))-)))))))) ( -33.30) >DroYak_CAF1 18381 102 - 1 AAUAAAAGAGGAAAAAGGAUUCUCAUUCCAUCUCCAAUUAUGGGGCAACCUCAGUGACCCACAUCUGCCCGUAGG-GAAAUGACCACAUUUCCCUGGAGGA-------------AU-- .........((((..((....))..)))).(((((.......(((((......(((...)))...)))))..(((-((((((....)))))))))))))).-------------..-- ( -30.70) >consensus AAUAAA________AAGGAUUCACAUUCCAACUCCCAUUCUGGGACAACCUCAGCGAUCCACAACUGCCUGGGGC_AAAAUGGGCCCAUUUCCCCAAGGGAAACAACGA_GGGAAUCC ................(((.......)))...((((...(((((.....)))))...............(((((...(((((....)))))))))).))))................. (-15.71 = -16.90 + 1.19)

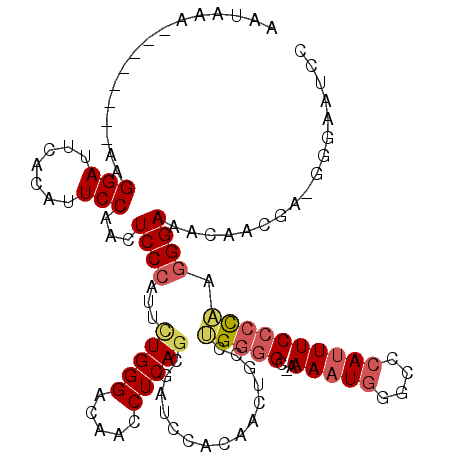
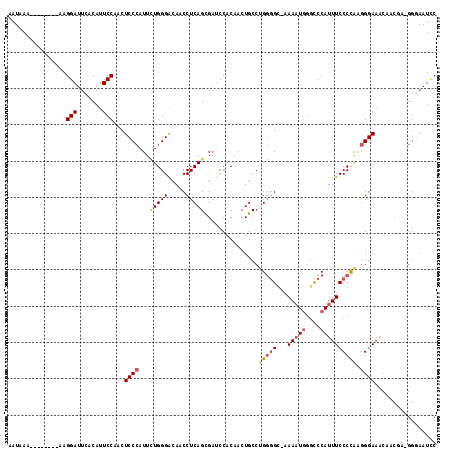
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:15 2006