
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,012,942 – 11,013,041 |
| Length | 99 |
| Max. P | 0.974876 |

| Location | 11,012,942 – 11,013,041 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 77.52 |
| Mean single sequence MFE | -24.08 |
| Consensus MFE | -12.85 |
| Energy contribution | -13.30 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952318 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 11012942 99 + 22224390 UAAAAAAAAAAAAAG-AAAAAACAACAAAAAUUGUCAAUUAAAGCCAUUAGGGCCAUUGGCAAUUAGUUGCCAGCCAGCCCCAUUAUGCGGAAAGUCAUU--- ...............-...........................((.....((((..(((((((....)))))))...))))......))...........--- ( -16.60) >DroPse_CAF1 4891 96 + 1 AGA-GCC-AUUUGCCCGUUGGCCGACAAAAAUUGUCAAUUAAAGCCAUUAAGGCCAACGGCAAUUAGUUGGCU----GC-CCAUUUUGUGGAAACUCAUUCCG ..(-(((-(.(((((.(((((((((((.....))))..((((.....)))))))))))))))).....)))))----..-.........((((.....)))). ( -33.10) >DroSec_CAF1 5837 83 + 1 AAAAAAA-------------GCCAACAAAAAUUGUCAAUUAAAGCCAUUAGGGCCAUUGGCAAUUAGUUGCCA----GCCCCAUUAUGCGGAAAGUCAUU--- .......-------------.((.(((.....)))........((.....((((...((((((....))))))----))))......)))).........--- ( -16.20) >DroSim_CAF1 909 84 + 1 AAAAAAA------------GGCCAACAAAAAUUGUCAAUUAAAGCCAUUAGGGCCAUUGGCAAUUAGUUGCCA----GCCCCAUUAUGCGGAAAGUCAUU--- .......------------(((.(((...((((((((((....(((.....))).)))))))))).)))))).----((........))...........--- ( -18.70) >DroYak_CAF1 903 94 + 1 AACAACC-CUGUGCU-GGCGGCCAACAGAAAUUGUCAAUUAAAGCCAUUAGGGCCAUUGGCAAUUAGUUGCCA----GCCCCAUUAUGCGGAAAGUCAUU--- .......-(((((((-(((((((....).((((((((((....(((.....))).)))))))))).)))))))----))........)))).........--- ( -26.70) >DroPer_CAF1 4870 96 + 1 AGA-GCC-AUUUGCCCGUUGGCCGACAAAAAUUGUCAAUUAAAGCCAUUAAGGCCAACGGCAAUUAGUUGGCU----GC-CCAUUAUGUGGAAACUCAUUCCG ..(-(((-(.(((((.(((((((((((.....))))..((((.....)))))))))))))))).....)))))----..-.....(((.(....).))).... ( -33.20) >consensus AAAAAAA_AU_UGC__G__GGCCAACAAAAAUUGUCAAUUAAAGCCAUUAGGGCCAUUGGCAAUUAGUUGCCA____GCCCCAUUAUGCGGAAAGUCAUU___ ......................((((...((((((((((....(((.....))).)))))))))).))))..........((.......))............ (-12.85 = -13.30 + 0.45)


| Location | 11,012,942 – 11,013,041 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 77.52 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -13.64 |
| Energy contribution | -14.28 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.49 |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.974876 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
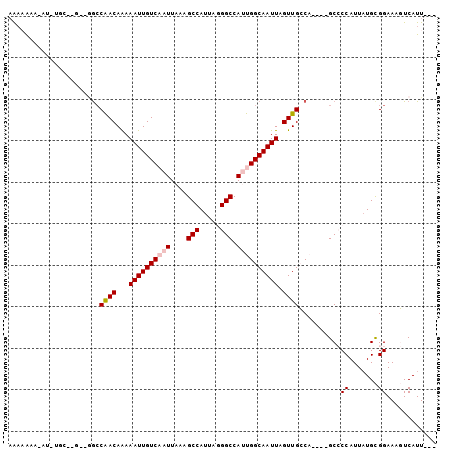
>X_DroMel_CAF1 11012942 99 - 22224390 ---AAUGACUUUCCGCAUAAUGGGGCUGGCUGGCAACUAAUUGCCAAUGGCCCUAAUGGCUUUAAUUGACAAUUUUUGUUGUUUUUU-CUUUUUUUUUUUUUA ---...........((....((((((((..((((((....)))))).))))))))...))...((..((((((....))))))..))-............... ( -24.70) >DroPse_CAF1 4891 96 - 1 CGGAAUGAGUUUCCACAAAAUGG-GC----AGCCAACUAAUUGCCGUUGGCCUUAAUGGCUUUAAUUGACAAUUUUUGUCGGCCAACGGGCAAAU-GGC-UCU .((((.....))))........(-(.----(((((.....((((((((((((((((.....))))..((((.....))))))))))).))))).)-)))-))) ( -32.50) >DroSec_CAF1 5837 83 - 1 ---AAUGACUUUCCGCAUAAUGGGGC----UGGCAACUAAUUGCCAAUGGCCCUAAUGGCUUUAAUUGACAAUUUUUGUUGGC-------------UUUUUUU ---..................(((((----..((((..(((((.((((((((.....))))...)))).))))).))))..))-------------))).... ( -21.00) >DroSim_CAF1 909 84 - 1 ---AAUGACUUUCCGCAUAAUGGGGC----UGGCAACUAAUUGCCAAUGGCCCUAAUGGCUUUAAUUGACAAUUUUUGUUGGCC------------UUUUUUU ---..................(((((----..((((..(((((.((((((((.....))))...)))).))))).))))..)))------------))..... ( -23.80) >DroYak_CAF1 903 94 - 1 ---AAUGACUUUCCGCAUAAUGGGGC----UGGCAACUAAUUGCCAAUGGCCCUAAUGGCUUUAAUUGACAAUUUCUGUUGGCCGCC-AGCACAG-GGUUGUU ---...(((..(((......((((((----..(((...(((((.((((((((.....))))...)))).)))))..)))..))).))-).....)-))..))) ( -31.50) >DroPer_CAF1 4870 96 - 1 CGGAAUGAGUUUCCACAUAAUGG-GC----AGCCAACUAAUUGCCGUUGGCCUUAAUGGCUUUAAUUGACAAUUUUUGUCGGCCAACGGGCAAAU-GGC-UCU .((((.....))))........(-(.----(((((.....((((((((((((((((.....))))..((((.....))))))))))).))))).)-)))-))) ( -32.50) >consensus ___AAUGACUUUCCGCAUAAUGGGGC____UGGCAACUAAUUGCCAAUGGCCCUAAUGGCUUUAAUUGACAAUUUUUGUUGGCC__C__GCA_AU_GGUUUUU ....................((((((....((((((....))))))...))))))..((((......((((.....))))))))................... (-13.64 = -14.28 + 0.64)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:53 2006