
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,999,502 – 10,999,622 |
| Length | 120 |
| Max. P | 0.860506 |

| Location | 10,999,502 – 10,999,622 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.89 |
| Mean single sequence MFE | -37.28 |
| Consensus MFE | -22.08 |
| Energy contribution | -24.32 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.563460 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10999502 120 + 22224390 AGUGAUUGUUGUGGGAACGGUAUUUCCCGCACCUGCUAGUGCCAAGUGUGACCAUACCCGGGUCACCAUUUGGUAUCGGUAUUUUUUGCAGUAUUUUUACACGAAUAAUUGGUGCGAAAG .........((((((((......))))))))..(((..(((((((((((((((.......))))).))))))))))..)))..((((((((((....))).(((....))).))))))). ( -41.40) >DroSim_CAF1 13875 120 + 1 AGUGAUUGUUGUGGGAACGUUAUUUCCCGCACCUGCUAGUGCCGAGUGUGACCAUACCCGGCUGGCCUUUUGGUAUCAGUAUUUUUUGCAGUAUUUCUACACGAAUAAUUGGUGCGAAAG .........((((((((......))))))))...((((..((((.((((....)))).))))))))..(((.((((((((..(((.((.((.....)).)).)))..)))))))).))). ( -34.50) >DroEre_CAF1 11096 113 + 1 AGG----GGUAAAGAAGCAGUAUUUUCCGCACCUGCUAGUGC--AGUGUGGCCAUACCCGGCUGGC-UAUUGGUAUCGGUAUUUUUUGCAGUAUUUUUGCACGAAUUAUUGGUGCGAAAG ...----(((.....(((.((((...((((((.(((....))--)))))))..))))...))).))-).((.(((((((((.(((.(((((.....))))).))).))))))))).)).. ( -35.70) >DroYak_CAF1 11918 119 + 1 AGUGACUGGUAUAGAAACAGUAUUUUCAGCACCUG-UAGGACAGAGUGUGACCAUACCAGUCUGGCCUUUUGGUAUCAGUAUUUUUGGCAGUAUUUUUGCACGAAUAAUUGGUGCGAAAG ...(((((((((.((((......)))).(((((((-.....))).))))....)))))))))......(((.((((((((...((((((((.....)))).))))..)))))))).))). ( -37.50) >consensus AGUGAUUGGUAUAGAAACAGUAUUUCCCGCACCUGCUAGUGCCGAGUGUGACCAUACCCGGCUGGCCUUUUGGUAUCAGUAUUUUUUGCAGUAUUUUUACACGAAUAAUUGGUGCGAAAG .......((((((((((......)))))))))).((((((.....((((....))))...))))))..(((.((((((((..(((.(((((.....))))).)))..)))))))).))). (-22.08 = -24.32 + 2.25)

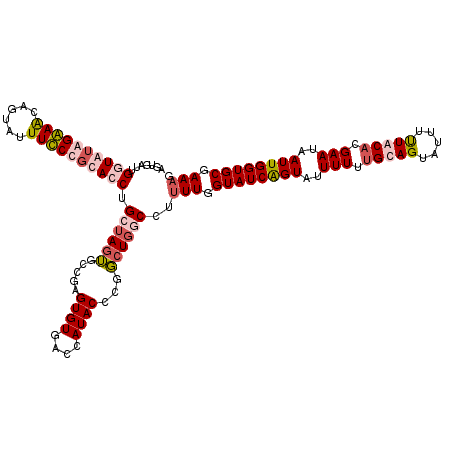
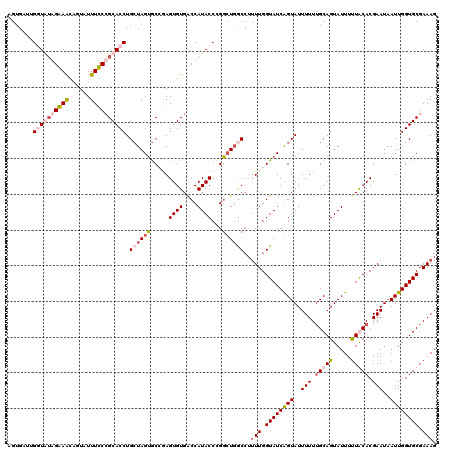
| Location | 10,999,502 – 10,999,622 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.89 |
| Mean single sequence MFE | -29.65 |
| Consensus MFE | -19.30 |
| Energy contribution | -19.80 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.860506 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10999502 120 - 22224390 CUUUCGCACCAAUUAUUCGUGUAAAAAUACUGCAAAAAAUACCGAUACCAAAUGGUGACCCGGGUAUGGUCACACUUGGCACUAGCAGGUGCGGGAAAUACCGUUCCCACAACAAUCACU .....(((((......(((.(((................))))))..((((...((((((.......))))))..))))........)))))(((((......)))))............ ( -30.89) >DroSim_CAF1 13875 120 - 1 CUUUCGCACCAAUUAUUCGUGUAGAAAUACUGCAAAAAAUACUGAUACCAAAAGGCCAGCCGGGUAUGGUCACACUCGGCACUAGCAGGUGCGGGAAAUAACGUUCCCACAACAAUCACU .....(((((..........(((....)))(((..............((....))...(((((((........)))))))....))))))))(((((......)))))............ ( -34.10) >DroEre_CAF1 11096 113 - 1 CUUUCGCACCAAUAAUUCGUGCAAAAAUACUGCAAAAAAUACCGAUACCAAUA-GCCAGCCGGGUAUGGCCACACU--GCACUAGCAGGUGCGGAAAAUACUGCUUCUUUACC----CCU .(((((((((.........((((.......))))...................-((((........)))).....(--((....)))))))))))).................----... ( -24.20) >DroYak_CAF1 11918 119 - 1 CUUUCGCACCAAUUAUUCGUGCAAAAAUACUGCCAAAAAUACUGAUACCAAAAGGCCAGACUGGUAUGGUCACACUCUGUCCUA-CAGGUGCUGAAAAUACUGUUUCUAUACCAGUCACU .....((((.........)))).........(((...................)))..(((((((((((((((((.(((.....-)))))).)))...........)))))))))))... ( -29.41) >consensus CUUUCGCACCAAUUAUUCGUGCAAAAAUACUGCAAAAAAUACCGAUACCAAAAGGCCAGCCGGGUAUGGUCACACUCGGCACUAGCAGGUGCGGAAAAUACCGUUCCCACAACAAUCACU .(((((((((.........((((.......))))......(((.(((((....((....)).)))))))).................)))))))))........................ (-19.30 = -19.80 + 0.50)


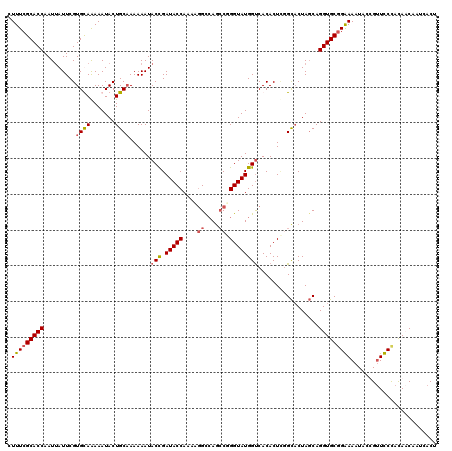
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:47 2006