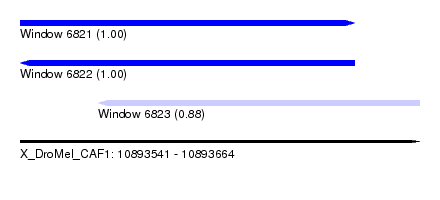
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,893,541 – 10,893,664 |
| Length | 123 |
| Max. P | 0.999901 |

| Location | 10,893,541 – 10,893,644 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 74.55 |
| Mean single sequence MFE | -29.10 |
| Consensus MFE | -16.51 |
| Energy contribution | -16.51 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.86 |
| Structure conservation index | 0.57 |
| SVM decision value | 3.46 |
| SVM RNA-class probability | 0.999250 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10893541 103 + 22224390 GGUGGGCAGUGAUGGGCGGGUCAACGUAAUCUGCCAUAAUCUUAUGGCAUCUUAUCCUUUCACAAAAUUUUGACCCAC-CCCACCCCACUCACUGAGCGAUGUC ((((((..(((..(((.(((((((.((((..(((((((....)))))))..))))..............))))))).)-))))))))))).............. ( -34.56) >DroSec_CAF1 47516 81 + 1 GGUGGGCAGUGAUGGGCGGG-------------CCAUAAUCCGA------AUUAUCCUUUCACAAAAA-UUGACCCAC-CCCACCC--AUCACUGAGCGACAUC .((.(.((((((((((.(((-------------.........((------(.......))).((....-.))......-))).)))--)))))))..).))... ( -26.80) >DroSim_CAF1 50673 81 + 1 GGUGGGCAGUGAUGGGCGGG-------------CCAUAAUCCGA------AUUAUCCUUUCACAAAAU-UUGACCCAU-CCCACCC--AUCACUGAGCGACAUC .((.(.((((((((((.(((-------------........(((------(((............)))-)))......-))).)))--)))))))..).))... ( -27.34) >DroEre_CAF1 24420 87 + 1 GGCGGGCAGUGAUGGGCGGG-CACU-----AUGCCAUGAUUAGA------AUUAUACUUUCG-AAAAC-CUGACCCAG-CCCACCC--AUCACUGAGCGAUAUC .((...((((((((((.(((-(.((-----(.........))).------.........(((-.....-.)))....)-))).)))--))))))).))...... ( -31.80) >DroYak_CAF1 48074 87 + 1 AGCGGGCAGUGAUGGGCGGGUCACC-------GUAAUAAUUGGA------AUUAAACUUUUC-AAAAC-AUGACCCAAUCCCACCC--AUCACUAAGUGAUUUG .......(((((((((.(((.....-------((.....(((((------(.......))))-))...-...)).....))).)))--)))))).......... ( -25.00) >consensus GGUGGGCAGUGAUGGGCGGG_CA_________GCCAUAAUCCGA______AUUAUCCUUUCACAAAAC_UUGACCCAC_CCCACCC__AUCACUGAGCGAUAUC .((...((((((((((.(((...........................................................))).)))..))))))).))...... (-16.51 = -16.51 + -0.00)


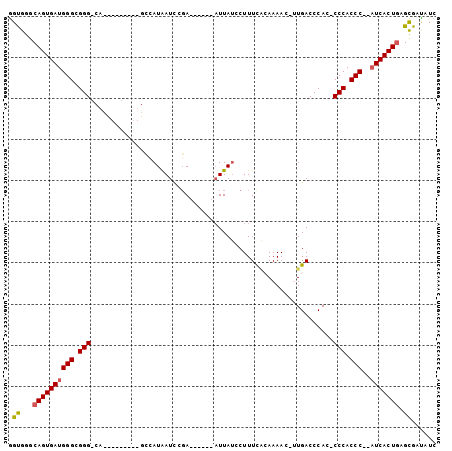
| Location | 10,893,541 – 10,893,644 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 74.55 |
| Mean single sequence MFE | -34.88 |
| Consensus MFE | -17.57 |
| Energy contribution | -17.61 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.51 |
| Structure conservation index | 0.50 |
| SVM decision value | 4.45 |
| SVM RNA-class probability | 0.999901 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10893541 103 - 22224390 GACAUCGCUCAGUGAGUGGGGUGGG-GUGGGUCAAAAUUUUGUGAAAGGAUAAGAUGCCAUAAGAUUAUGGCAGAUUACGUUGACCCGCCCAUCACUGCCCACC ....((((...))))((((((((((-(((((((((.(((((((......)))))))((((((....))))))........)))))))))))..)))..))))). ( -42.10) >DroSec_CAF1 47516 81 - 1 GAUGUCGCUCAGUGAU--GGGUGGG-GUGGGUCAA-UUUUUGUGAAAGGAUAAU------UCGGAUUAUGG-------------CCCGCCCAUCACUGCCCACC ......(..(((((((--(((((((-...((((..-...((((......)))).------...))))....-------------))))))))))))))..)... ( -31.00) >DroSim_CAF1 50673 81 - 1 GAUGUCGCUCAGUGAU--GGGUGGG-AUGGGUCAA-AUUUUGUGAAAGGAUAAU------UCGGAUUAUGG-------------CCCGCCCAUCACUGCCCACC ......(..(((((((--(((((((-...((((..-...((((......)))).------...))))....-------------))))))))))))))..)... ( -32.10) >DroEre_CAF1 24420 87 - 1 GAUAUCGCUCAGUGAU--GGGUGGG-CUGGGUCAG-GUUUU-CGAAAGUAUAAU------UCUAAUCAUGGCAU-----AGUG-CCCGCCCAUCACUGCCCGCC .....((..(((((((--(((((((-(...(((((-(((..-.(((.......)------)).)))).))))..-----...)-))))))))))))))..)).. ( -38.20) >DroYak_CAF1 48074 87 - 1 CAAAUCACUUAGUGAU--GGGUGGGAUUGGGUCAU-GUUUU-GAAAAGUUUAAU------UCCAAUUAUUAC-------GGUGACCCGCCCAUCACUGCCCGCU .........(((((((--(((((((((((((...(-(..((-....))..))..------))))))).....-------(....))))))))))))))...... ( -31.00) >consensus GAUAUCGCUCAGUGAU__GGGUGGG_GUGGGUCAA_AUUUUGUGAAAGGAUAAU______UCGGAUUAUGGC_________UG_CCCGCCCAUCACUGCCCACC .........(((((((..(((((((...........................................................))))))))))))))...... (-17.57 = -17.61 + 0.04)

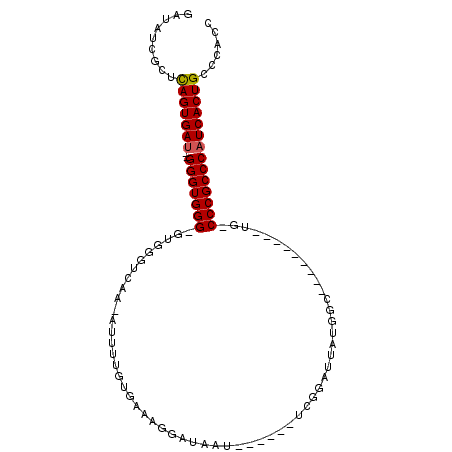
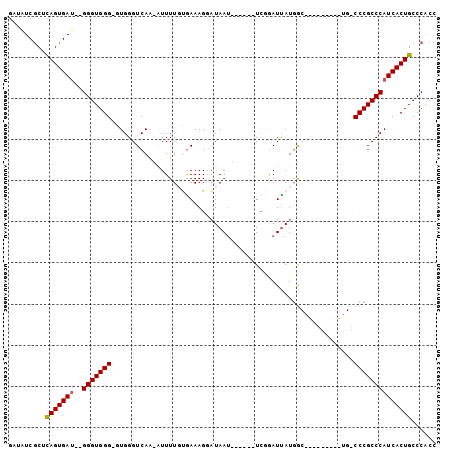
| Location | 10,893,565 – 10,893,664 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 74.14 |
| Mean single sequence MFE | -24.92 |
| Consensus MFE | -15.46 |
| Energy contribution | -15.14 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.884299 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10893565 99 - 22224390 UACAGCCCAUCUGGCCCAAGGACAUCGCUCAGUGAGUGGGGUGGG-GUGGGUCAAAAUUUUGUGAAAGGAUAAGAUGCCAUAAGAUUAUGGCAGAUUACG ....((((((((.((((....((.((((...)))))).)))).))-))))))..................(((..(((((((....)))))))..))).. ( -34.00) >DroSec_CAF1 47536 81 - 1 UACAGCCCAAUUGGCCCAAGGAUGUCGCUCAGUGAU--GGGUGGG-GUGGGUCAA-UUUUUGUGAAAGGAUAAU------UCGGAUUAUGG--------- (((((...((((((((((....(.(((((((....)--)))))).-)))))))))-)).)))))..........------...........--------- ( -23.70) >DroSim_CAF1 50693 81 - 1 UACAGCCCAAUUGGCCCAAGGAUGUCGCUCAGUGAU--GGGUGGG-AUGGGUCAA-AUUUUGUGAAAGGAUAAU------UCGGAUUAUGG--------- ......((..((((((((....(.(((((((....)--)))))).-)))))))))-(((((.....)))))...------..)).......--------- ( -22.10) >DroEre_CAF1 24443 83 - 1 CACAGCC-UUUUGGCCCAAGGAUAUCGCUCAGUGAU--GGGUGGG-CUGGGUCAG-GUUUU-CGAAAGUAUAAU------UCUAAUCAUGGCAU-----A ....(((-.(((((((((..(...(((((((....)--)))))).-)))))))))-)....-(....)......------.........)))..-----. ( -25.30) >DroYak_CAF1 48098 83 - 1 UACAUCCAUUUUGGCCCAAGCAAAUCACUUAGUGAU--GGGUGGGAUUGGGUCAU-GUUUU-GAAAAGUUUAAU------UCCAAUUAUUAC-------G ...........((((((((.....(((((((....)--))))))..)))))))).-.....-............------............-------. ( -19.50) >consensus UACAGCCCAUUUGGCCCAAGGAUAUCGCUCAGUGAU__GGGUGGG_GUGGGUCAA_AUUUUGUGAAAGGAUAAU______UCGGAUUAUGGC________ ...........(((((((......((((((........))))))...))))))).............................................. (-15.46 = -15.14 + -0.32)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:42 2006