
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,856,469 – 10,856,588 |
| Length | 119 |
| Max. P | 0.887899 |

| Location | 10,856,469 – 10,856,588 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.22 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -19.20 |
| Energy contribution | -19.95 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.887899 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
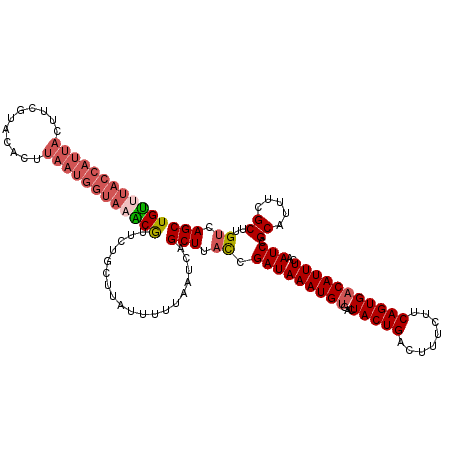
>X_DroMel_CAF1 10856469 119 + 22224390 GCUGACAAGCGAAAUGCGGAUUGAAAUGUCACUGAAGAAAGUCAGUAGUCACAUUUAUCGGUAAGCUGAUUAA-AAUAAACAGAAUGCAUACCAUUAAGUGUAAGAAGUAAUGGUAAACA (((....)))....((((..(((((((((.((((..(.....)..)))).))))))(((((....)))))...-......)))..))))((((((((..(.....)..)))))))).... ( -26.60) >DroSec_CAF1 10309 120 + 1 GCUGACAAGCGAAAUGCGGAUUGAAAUGUCACUGAAGAAAGUCAGUAGUCUCAUUUAUCGGUAAGCUGAUUAAAAAUAAGCAAAACGUUUACCAUUAAGUGUACGAAGUAAUGGUAAACA (((....)))....(((..(((.(((((.((((((......))))).)...)))))(((((....)))))....)))..)))....(((((((((((..(.....)..))))))))))). ( -29.70) >DroSim_CAF1 12231 120 + 1 GCUGACAAGCAAAAUGCGGAUUGAAAUGUCACUGAAGAAAGUCAGUAGUCACAUUUAUCGGUAAGCUGAUUAAAAAUAAGCAGAACGUUUACCAUUAAGUGUACGAAGUAAUGGUAAACA (((....)))....(((..(((.((((((.((((..(.....)..)))).))))))(((((....)))))....)))..)))....(((((((((((..(.....)..))))))))))). ( -30.50) >DroYak_CAF1 10447 110 + 1 GCUGACAAGCGAAAUGCGGAUUGAAAUGUCACUGCAGAAAGUCAGUAGUCACAUUUAUCGAAAAGCUGAUUCAAAAU-AGCAGAACGCUUACAACUAAGUGCUAAAA---------CGCA (((....)))....(((((((..((((((.(((((.(.....).))))).))))))))).....((((........)-)))....((((((....))))))......---------)))) ( -23.90) >consensus GCUGACAAGCGAAAUGCGGAUUGAAAUGUCACUGAAGAAAGUCAGUAGUCACAUUUAUCGGUAAGCUGAUUAAAAAUAAGCAGAACGCUUACCAUUAAGUGUAAGAAGUAAUGGUAAACA ((((((..((.....)).(((......)))..........)))))).....((((((..(((((((....................)))))))..))))))................... (-19.20 = -19.95 + 0.75)



| Location | 10,856,469 – 10,856,588 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.22 |
| Mean single sequence MFE | -23.30 |
| Consensus MFE | -15.09 |
| Energy contribution | -17.27 |
| Covariance contribution | 2.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.551557 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10856469 119 - 22224390 UGUUUACCAUUACUUCUUACACUUAAUGGUAUGCAUUCUGUUUAUU-UUAAUCAGCUUACCGAUAAAUGUGACUACUGACUUUCUUCAGUGACAUUUCAAUCCGCAUUUCGCUUGUCAGC (((.((((((((...........)))))))).)))...........-.......(((.((.(((((((((...((((((......))))))))))))..))).((.....))..)).))) ( -20.40) >DroSec_CAF1 10309 120 - 1 UGUUUACCAUUACUUCGUACACUUAAUGGUAAACGUUUUGCUUAUUUUUAAUCAGCUUACCGAUAAAUGAGACUACUGACUUUCUUCAGUGACAUUUCAAUCCGCAUUUCGCUUGUCAGC .(((((((((((...........)))))))))))((...(((.((.....)).)))..)).(((((.(((((.((((((......))))))...)))))....((.....)))))))... ( -24.60) >DroSim_CAF1 12231 120 - 1 UGUUUACCAUUACUUCGUACACUUAAUGGUAAACGUUCUGCUUAUUUUUAAUCAGCUUACCGAUAAAUGUGACUACUGACUUUCUUCAGUGACAUUUCAAUCCGCAUUUUGCUUGUCAGC .(((((((((((...........)))))))))))....................(((.((.(((((((((...((((((......))))))))))))..))).((.....))..)).))) ( -23.40) >DroYak_CAF1 10447 110 - 1 UGCG---------UUUUAGCACUUAGUUGUAAGCGUUCUGCU-AUUUUGAAUCAGCUUUUCGAUAAAUGUGACUACUGACUUUCUGCAGUGACAUUUCAAUCCGCAUUUCGCUUGUCAGC (((.---------.....)))....((((((((((...(((.-...(((((.......))))).((((((.(((.(.(.....).).))).))))))......)))...)))))).)))) ( -24.80) >consensus UGUUUACCAUUACUUCGUACACUUAAUGGUAAACGUUCUGCUUAUUUUUAAUCAGCUUACCGAUAAAUGUGACUACUGACUUUCUUCAGUGACAUUUCAAUCCGCAUUUCGCUUGUCAGC ((((((((((((...........))))))))))))...................(((.((.(((((((((...(((((........)))))))))))..))).((.....))..)).))) (-15.09 = -17.27 + 2.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:00 2006