
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,827,697 – 10,827,794 |
| Length | 97 |
| Max. P | 0.990384 |

| Location | 10,827,697 – 10,827,794 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 74.62 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -17.96 |
| Energy contribution | -17.85 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
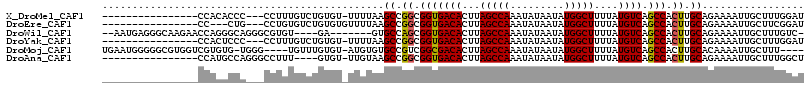
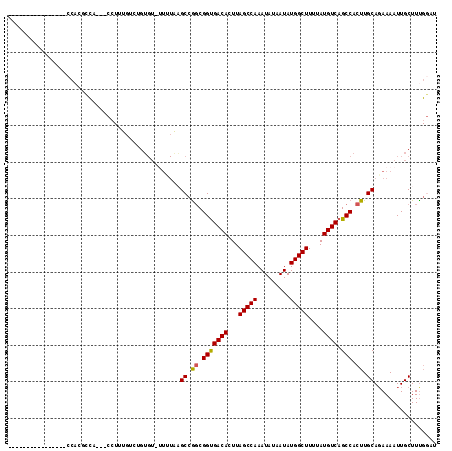
>X_DroMel_CAF1 10827697 97 + 22224390 ----------------CCACACCC---CCUUUGUCUGUGU-UUUUAAGCCGGCGGUGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCAGAAAAUUGCUUUGGAU ----------------........---((...((....((-((((..((.((.(((((((...(((((.........)))))....))))).)).)).)).)))))).))...)).. ( -21.50) >DroEre_CAF1 51185 95 + 1 ----------------CC---CUG---CCUGUGUCUGUGUGUUUUAAGCCGGCGGUGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCAGAAAAUUGCUUCGGAU ----------------..---...---.....(((((...(((((..((.((.(((((((...(((((.........)))))....))))).)).)).))..))))).....))))) ( -24.20) >DroWil_CAF1 42008 103 + 1 --AAUGAGGGCAAGAACCAGGGCAGGGCGUGU----GA-------GUGCCAGCGGUGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCAGAAAAUUGCUUUGUC- --.....((.......))..(((((((((...----..-------.(((.((.(((((((...(((((.........)))))....))))).)).)).)))......)))))))))- ( -31.20) >DroYak_CAF1 46960 97 + 1 ----------------CCACUCCC---CCUUUGUCUGUGU-UUUUAAGCCGGCGGUGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCAGAAAAUUGCUUUGGAU ----------------........---((...((....((-((((..((.((.(((((((...(((((.........)))))....))))).)).)).)).)))))).))...)).. ( -21.50) >DroMoj_CAF1 50728 107 + 1 UGAAUGGGGGCGUGGUCGUGUG-UGGG----UGUUUGUGU-AUGUGUGCCGUCGGCGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCACAAAAUUGCUUU---- ...........(..((..((((-..((----((.((((((-(.(((((((...))).))))..(((((.........)))))..)))).))).))))..))))..))..)...---- ( -32.40) >DroAna_CAF1 51944 96 + 1 ----------------CCAUGCCAGGGCCUUU----GUGU-UUGUAAGCCGGCGGUGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCAGAAAAUUGCUUUGGCU ----------------....((((((((..((----...(-(((((((..(((..(((((...(((((.........)))))....)))))))).)))))))).))..)))))))). ( -37.00) >consensus ________________CCACGCCA___CCUUUGUCUGUGU_UUUUAAGCCGGCGGUGACACUUAGCCAAAUAUAAUAUGGCUUUUAUGUCAGCCACUUGCAGAAAAUUGCUUUGGAU ...............................................((.((.(((((((...(((((.........)))))....)))).))).)).))................. (-17.96 = -17.85 + -0.11)
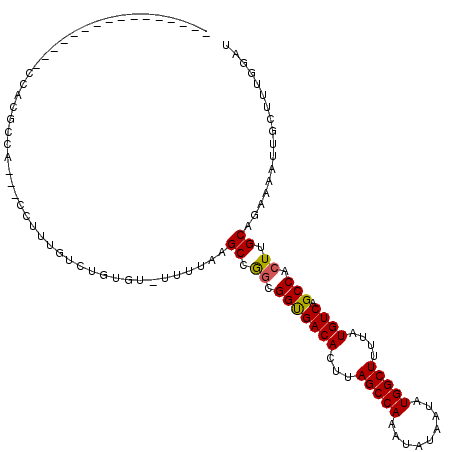

| Location | 10,827,697 – 10,827,794 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 74.62 |
| Mean single sequence MFE | -27.48 |
| Consensus MFE | -20.92 |
| Energy contribution | -20.95 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.76 |
| SVM decision value | 2.21 |
| SVM RNA-class probability | 0.990384 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
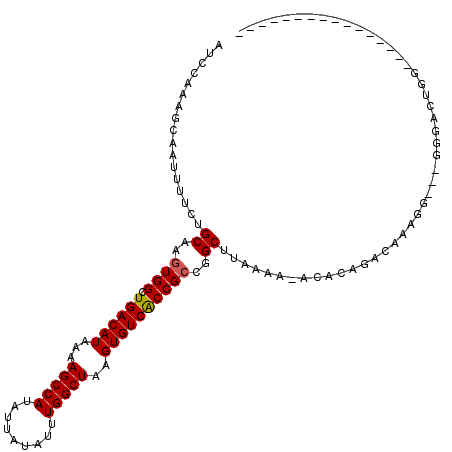
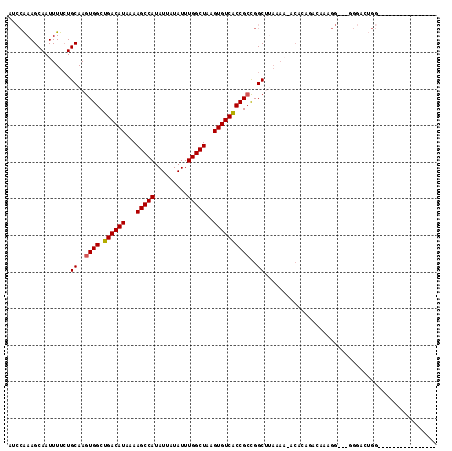
>X_DroMel_CAF1 10827697 97 - 22224390 AUCCAAAGCAAUUUUCUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCACCGCCGGCUUAAAA-ACACAGACAAAGG---GGGUGUGG---------------- ...........((((..((..((((.((((((...(((((.........)))))..))))))))))..))..))))-.((((..(.....---)..)))).---------------- ( -27.90) >DroEre_CAF1 51185 95 - 1 AUCCGAAGCAAUUUUCUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCACCGCCGGCUUAAAACACACAGACACAGG---CAG---GG---------------- ..((...((.....((((...((((.((((((...(((((.........)))))..)))))))))).(........)...)))).....)---).)---).---------------- ( -24.80) >DroWil_CAF1 42008 103 - 1 -GACAAAGCAAUUUUCUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCACCGCUGGCAC-------UC----ACACGCCCUGCCCUGGUUCUUGCCCUCAUU-- -......((((...((.((((((((.((((((...(((((.........)))))..)))))))))))(((..-------..----....))).)))...))...)))).......-- ( -29.20) >DroYak_CAF1 46960 97 - 1 AUCCAAAGCAAUUUUCUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCACCGCCGGCUUAAAA-ACACAGACAAAGG---GGGAGUGG---------------- .(((...(((......)))(((((((((((((...(((((.........)))))..))))))..))).))))....-.............---.)))....---------------- ( -25.00) >DroMoj_CAF1 50728 107 - 1 ----AAAGCAAUUUUGUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCGCCGACGGCACACAU-ACACAAACA----CCCA-CACACGACCACGCCCCCAUUCA ----..........(((((...((((.(((((...(((((.........)))))..)))))))))...)))))...-.........----....-...................... ( -26.20) >DroAna_CAF1 51944 96 - 1 AGCCAAAGCAAUUUUCUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCACCGCCGGCUUACAA-ACAC----AAAGGCCCUGGCAUGG---------------- .((((..(((......)))..((((.((((((...(((((.........)))))..)))))))))).(((((....-....----..))))).))))....---------------- ( -31.80) >consensus AUCCAAAGCAAUUUUCUGCAAGUGGCUGACAUAAAAGCCAUAUUAUAUUUGGCUAAGUGUCACCGCCGGCUUAAAA_ACACAGACAAAGG___GGGACUGG________________ .................((..((((.((((((...(((((.........)))))..))))))))))..))............................................... (-20.92 = -20.95 + 0.03)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:16:19 2006