
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,782,453 – 10,782,587 |
| Length | 134 |
| Max. P | 0.974223 |

| Location | 10,782,453 – 10,782,547 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 95.97 |
| Mean single sequence MFE | -25.00 |
| Consensus MFE | -21.60 |
| Energy contribution | -21.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.548222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10782453 94 + 22224390 UUCAUUUGAUGUUUCUGUUGUUUUGGGCAUAAAUAUUUAUGUCUGUGU------GUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGA ((((((((((..............((((((((....))))))))(((.------((((((.....))))..)))))..))))))))))............ ( -23.20) >DroSec_CAF1 31927 98 + 1 UUCAUUUGAUGUUUCUGUUGUUUUGGGCAUAAAUAUUUAUGUCUGUGUGU--GUGUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGA ((((((((((..............((((((((....)))))))).((((.--(((.((((.....)))).))))))).))))))))))............ ( -25.90) >DroSim_CAF1 30667 100 + 1 UUCAUUUGAUGUUUCUGUUGUUUUGGGCAUAAAUAUUUAUGUCUGUGUGUGUGUGUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGA ((((((((((..............((((((((....))))))))...((((.(((.((((.....)))).))))))).))))))))))............ ( -25.90) >consensus UUCAUUUGAUGUUUCUGUUGUUUUGGGCAUAAAUAUUUAUGUCUGUGUGU__GUGUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGA ((((((((((..............((((((((....)))))))).............(((....(((....)))))).))))))))))............ (-21.60 = -21.60 + -0.00)

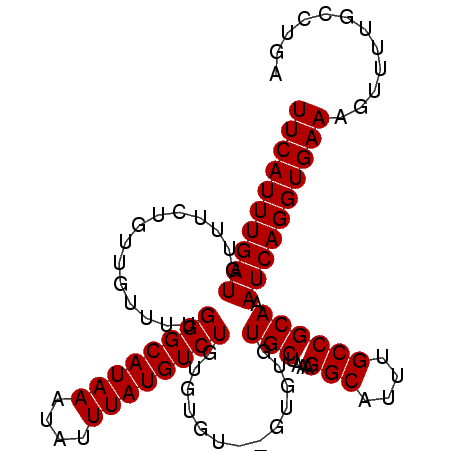

| Location | 10,782,493 – 10,782,587 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 93.22 |
| Mean single sequence MFE | -27.93 |
| Consensus MFE | -24.90 |
| Energy contribution | -24.90 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.49 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.73 |
| SVM RNA-class probability | 0.974223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10782493 94 + 22224390 GUCUGUGU------GUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGAGAAAAGGCAAAGACAAAACAAAAACAACAACAGCAACAUC (.((((.(------((.(((.....)))((((((.....(((((..(.....)..))))).....))))))............)))...)))))...... ( -26.70) >DroSec_CAF1 31967 95 + 1 GUCUGUGUGU--GUGUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGAGAAAAGGCAAAGACAAAACAAAAACAACAAC---AACAGC ..((((((.(--((.(((((.....)).((((((.....(((((..(.....)..))))).....))))))......)))...)))...))---.)))). ( -28.10) >DroSim_CAF1 30707 97 + 1 GUCUGUGUGUGUGUGUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGAGAAAAGGCAAAGACAAAACAAAAACAACAAC---AACAGC ..((((.(((.(((.(((((.....)).((((((.....(((((..(.....)..))))).....))))))......)))...)))...))---))))). ( -29.00) >consensus GUCUGUGUGU__GUGUGUGCUAAAGGCAUUUGCCGCAAAUCAGGUGAAAGUUUUGCCUGAGAAAAGGCAAAGACAAAACAAAAACAACAAC___AACAGC ((((............((((.....))))(((((.....(((((..(.....)..))))).....))))))))).......................... (-24.90 = -24.90 + 0.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:16:04 2006