
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,750,232 – 10,750,326 |
| Length | 94 |
| Max. P | 0.984582 |

| Location | 10,750,232 – 10,750,326 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 92.06 |
| Mean single sequence MFE | -21.60 |
| Consensus MFE | -17.31 |
| Energy contribution | -17.50 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.584156 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10750232 94 + 22224390 ACCUUUGCUUCUGU------UUCUGCUUCUUCCGCUACCUUUCUUCGUCAUUCUGCUCGGUUUUUUUGCGCAUGCGCCACCGAGACAGCGAAACAGCACA ..........((((------(((.((.......)).................(((((((((......(((....))).)))))).))).))))))).... ( -22.00) >DroSec_CAF1 26147 93 + 1 ACCUUUGCUUCUGU------UUCUGCUUCUUCCGCUACCUUUCUUCGUCAUUCUGCUCGGUUU-UUUGCGCAUGCGCCACCGAGACAGCGAAACAGCACA ..........((((------(((.((.......)).................(((((((((..-...(((....))).)))))).))).))))))).... ( -22.70) >DroEre_CAF1 29299 99 + 1 ACCUUUGCUUCUGUUUUUGUUUCUGUUUCUUCCGCUACCUUUCUUCGUCAUUCUGCUCGGUUU-UUUGCGCAUGCGCCAACGAGACAGCGAAGUAGCACA .................................(((((.....((((.....(((((((....-...(((....)))...)))).))))))))))))... ( -18.10) >DroYak_CAF1 26945 92 + 1 ACCUUUGCUUCUGC-------UCUGCUUCUUCCGCUACCUUUCUUCGUUAUUCUGCUCGGUUU-UUUGCGCAUGCGCCGCCGAGACAGCGAAGCAGCACA ......((....))-------.(((((((.......((........))....(((((((((..-...(((....))).)))))).))).))))))).... ( -23.60) >consensus ACCUUUGCUUCUGU______UUCUGCUUCUUCCGCUACCUUUCUUCGUCAUUCUGCUCGGUUU_UUUGCGCAUGCGCCACCGAGACAGCGAAACAGCACA ....(((((((.............((.......)).................(((((((((......(((....))).)))))).))).))))))).... (-17.31 = -17.50 + 0.19)


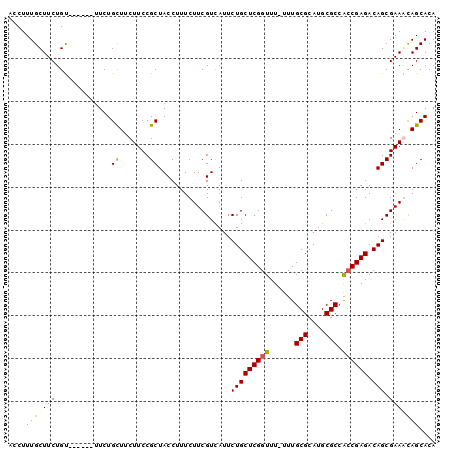
| Location | 10,750,232 – 10,750,326 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 92.06 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -24.61 |
| Energy contribution | -24.68 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.62 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.984582 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10750232 94 - 22224390 UGUGCUGUUUCGCUGUCUCGGUGGCGCAUGCGCAAAAAAACCGAGCAGAAUGACGAAGAAAGGUAGCGGAAGAAGCAGAA------ACAGAAGCAAAGGU (((.(((((((.(((.((((((.(((....)))......)))))))))..............((..(....)..)).)))------))))..)))..... ( -30.30) >DroSec_CAF1 26147 93 - 1 UGUGCUGUUUCGCUGUCUCGGUGGCGCAUGCGCAAA-AAACCGAGCAGAAUGACGAAGAAAGGUAGCGGAAGAAGCAGAA------ACAGAAGCAAAGGU (((.(((((((.(((.((((((.(((....)))...-..)))))))))..............((..(....)..)).)))------))))..)))..... ( -31.00) >DroEre_CAF1 29299 99 - 1 UGUGCUACUUCGCUGUCUCGUUGGCGCAUGCGCAAA-AAACCGAGCAGAAUGACGAAGAAAGGUAGCGGAAGAAACAGAAACAAAAACAGAAGCAAAGGU ..((((.((((((((.((((((.(((....)))...-.)).))))))).....)))))....((..(....)..))...............))))..... ( -23.00) >DroYak_CAF1 26945 92 - 1 UGUGCUGCUUCGCUGUCUCGGCGGCGCAUGCGCAAA-AAACCGAGCAGAAUAACGAAGAAAGGUAGCGGAAGAAGCAGA-------GCAGAAGCAAAGGU (((.(((((((.(((.(((((..(((....)))...-...)))))))).....((...........))...))))))).-------)))........... ( -27.70) >consensus UGUGCUGCUUCGCUGUCUCGGUGGCGCAUGCGCAAA_AAACCGAGCAGAAUGACGAAGAAAGGUAGCGGAAGAAGCAGAA______ACAGAAGCAAAGGU ..((((.((((((((.((((((.(((....)))......))))))))).....)))))....((..(....)..))...............))))..... (-24.61 = -24.68 + 0.06)

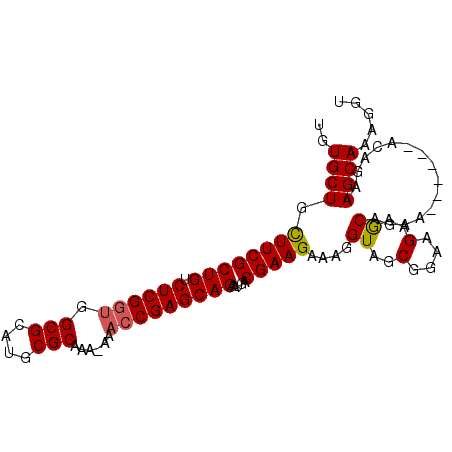

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:15:13 2006