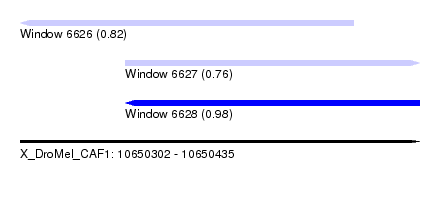
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,650,302 – 10,650,435 |
| Length | 133 |
| Max. P | 0.984516 |

| Location | 10,650,302 – 10,650,413 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 92.68 |
| Mean single sequence MFE | -20.64 |
| Consensus MFE | -17.02 |
| Energy contribution | -16.98 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.819737 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10650302 111 - 22224390 UAAACACUAAAUACUUUA-AGCGGCUCAAAUACCACACUUUUGAACCACUGUGCGCAUAUGUGUACUUCAAUUUCAAUCAUGCCC-----AGCAAAAGGUGCGAAUCCAUCAAAGUC ............(((((.-.((((.(((((.........))))).))...((((((....))))))...................-----.))....((((......))))))))). ( -18.80) >DroSec_CAF1 7756 111 - 1 UAAACACUAAAUACUUUA-AGCGGCUCAAAUACCACACUUUUGAACCACUGUGCGCACAUGUGUACUUCAAUUUCAAUCAUGCCC-----AGCAAAAGGUGCGAAUCCAUCAAAGUC ............(((((.-.((((.(((((.........))))).))...((((((....))))))...................-----.))....((((......))))))))). ( -18.80) >DroSim_CAF1 6044 111 - 1 UAAACACUAAAUACUUUA-AGCGGUUCAAAUACCACACUUUCGAACCACUGUGCGCACAUGUGUACUUCAAUUUCAAUCAUGCCC-----AGCAAAAGGUGCGAAUCCAUCAAAGUC ............(((((.-.(.(((......))).)...((((.(((...((((((....))))))..............(((..-----.)))...))).))))......))))). ( -20.10) >DroEre_CAF1 7810 112 - 1 UAAACACCAAAUACUUUAAAGUGGCUCAAAUACUACACUUUUGACCCACUGUGCGCACAUGUGUACUUCACAUUCAAUCAUUCAC-----AGCAAAAGGUGCGAAUCCAUCAAAGUC ....((((...........(((((.(((((.........))))).)))))((((((....))))))...................-----.......))))................ ( -22.90) >DroYak_CAF1 8085 116 - 1 UAAACACUAAAUACUUUA-AGUGGCUCAAACACUAUACUUUUGAUCCACUGUGCGCACAUGUGUACUUCAAUUUCAAUCAUGCCCAAGUGAGCAAAAGGUGCGAAUCCAUCAAAGUC ............(((((.-(((((.(((((.........))))).)))))((((((....))))))......(((..((((......))))(((.....))))))......))))). ( -22.60) >consensus UAAACACUAAAUACUUUA_AGCGGCUCAAAUACCACACUUUUGAACCACUGUGCGCACAUGUGUACUUCAAUUUCAAUCAUGCCC_____AGCAAAAGGUGCGAAUCCAUCAAAGUC ............(((((...((((.(((((.........))))).))...((((((....)))))).........................))....((((......))))))))). (-17.02 = -16.98 + -0.04)


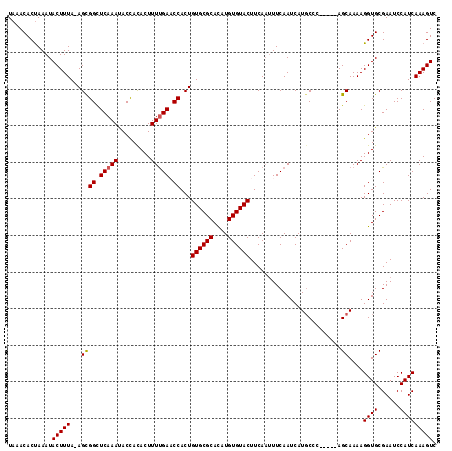
| Location | 10,650,337 – 10,650,435 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 87.22 |
| Mean single sequence MFE | -23.96 |
| Consensus MFE | -20.30 |
| Energy contribution | -20.66 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.762084 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10650337 98 + 22224390 UUGAAAUUGAAGUACACAUAUGCGCACAGUGGUUCAAAAGUGUGGUAUUUGAGCCGCU-UAAAGUAUUUAGUGUUUACCAUGGAAAUGUGGGUAUUUU-------U ........(((((((((((((((....(((((((((((.........)))))))))))-....))))...))))...((((......)))))))))))-------. ( -25.20) >DroSec_CAF1 7791 105 + 1 UUGAAAUUGAAGUACACAUGUGCGCACAGUGGUUCAAAAGUGUGGUAUUUGAGCCGCU-UAAAGUAUUUAGUGUUUACCAUGGAAAUGUGGGGAUUUUAUUAUAUU .(((((((.....((((..((((....(((((((((((.........)))))))))))-....))))...))))...((((......)))).)))))))....... ( -26.30) >DroSim_CAF1 6079 105 + 1 UUGAAAUUGAAGUACACAUGUGCGCACAGUGGUUCGAAAGUGUGGUAUUUGAACCGCU-UAAAGUAUUUAGUGUUUACCAUGGAAAUGUGGGGAUUUUAUUAUUUU .(((((((.....((((..((((....(((((((((((.........)))))))))))-....))))...))))...((((......)))).)))))))....... ( -26.00) >DroEre_CAF1 7845 96 + 1 UUGAAUGUGAAGUACACAUGUGCGCACAGUGGGUCAAAAGUGUAGUAUUUGAGCCACUUUAAAGUAUUUGGUGUUUACCAUGGAAAUG---AGAAUUU-------U .((((..(.((((((...(((....)))((((.(((((.........))))).))))......)))))).)..)))).(((....)))---.......-------. ( -22.20) >DroYak_CAF1 8125 98 + 1 UUGAAAUUGAAGUACACAUGUGCGCACAGUGGAUCAAAAGUAUAGUGUUUGAGCCACU-UAAAGUAUUUAGUGUUUACCAUGGUAAUGUGGGGAAUUU-------A ...(((((.....((((..((((....(((((.(((((.........))))).)))))-....))))...))))...((((......))))..)))))-------. ( -20.10) >consensus UUGAAAUUGAAGUACACAUGUGCGCACAGUGGUUCAAAAGUGUGGUAUUUGAGCCGCU_UAAAGUAUUUAGUGUUUACCAUGGAAAUGUGGGGAUUUU_______U .............((((..((((....(((((((((((.........))))))))))).....))))...))))...((((......))))............... (-20.30 = -20.66 + 0.36)


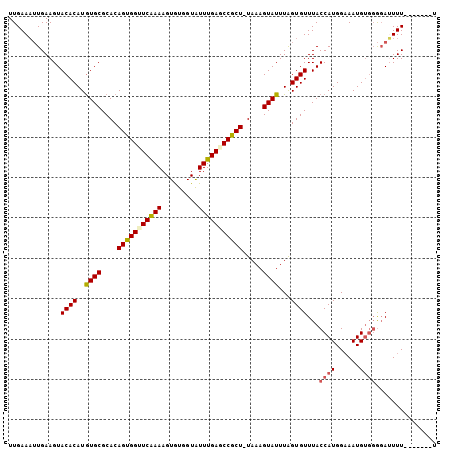
| Location | 10,650,337 – 10,650,435 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 87.22 |
| Mean single sequence MFE | -16.34 |
| Consensus MFE | -12.88 |
| Energy contribution | -12.92 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.984516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10650337 98 - 22224390 A-------AAAAUACCCACAUUUCCAUGGUAAACACUAAAUACUUUA-AGCGGCUCAAAUACCACACUUUUGAACCACUGUGCGCAUAUGUGUACUUCAAUUUCAA .-------....((((...........))))................-((.((.(((((.........))))).)).))((((((....))))))........... ( -14.80) >DroSec_CAF1 7791 105 - 1 AAUAUAAUAAAAUCCCCACAUUUCCAUGGUAAACACUAAAUACUUUA-AGCGGCUCAAAUACCACACUUUUGAACCACUGUGCGCACAUGUGUACUUCAAUUUCAA ...............(((........)))..................-((.((.(((((.........))))).)).))((((((....))))))........... ( -14.00) >DroSim_CAF1 6079 105 - 1 AAAAUAAUAAAAUCCCCACAUUUCCAUGGUAAACACUAAAUACUUUA-AGCGGUUCAAAUACCACACUUUCGAACCACUGUGCGCACAUGUGUACUUCAAUUUCAA ...............(((........)))..................-((.((((((((........))).))))).))((((((....))))))........... ( -14.40) >DroEre_CAF1 7845 96 - 1 A-------AAAUUCU---CAUUUCCAUGGUAAACACCAAAUACUUUAAAGUGGCUCAAAUACUACACUUUUGACCCACUGUGCGCACAUGUGUACUUCACAUUCAA .-------.......---........((((....))))..........(((((.(((((.........))))).)))))((((((....))))))........... ( -20.00) >DroYak_CAF1 8125 98 - 1 U-------AAAUUCCCCACAUUACCAUGGUAAACACUAAAUACUUUA-AGUGGCUCAAACACUAUACUUUUGAUCCACUGUGCGCACAUGUGUACUUCAAUUUCAA .-------............((((....))))...............-(((((.(((((.........))))).)))))((((((....))))))........... ( -18.50) >consensus A_______AAAAUCCCCACAUUUCCAUGGUAAACACUAAAUACUUUA_AGCGGCUCAAAUACCACACUUUUGAACCACUGUGCGCACAUGUGUACUUCAAUUUCAA ..........................((((....))))..........((.((.(((((.........))))).)).))((((((....))))))........... (-12.88 = -12.92 + 0.04)


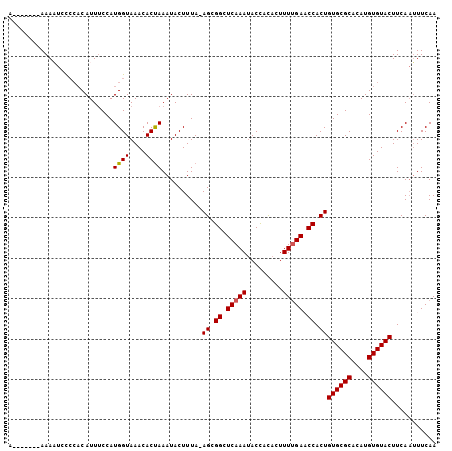
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:14:42 2006