
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,518,757 – 10,518,893 |
| Length | 136 |
| Max. P | 0.683314 |

| Location | 10,518,757 – 10,518,858 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 82.64 |
| Mean single sequence MFE | -17.75 |
| Consensus MFE | -12.30 |
| Energy contribution | -12.12 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.621125 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10518757 101 + 22224390 GUGGUGACUAAUUGUCGAUAAACAUGCUAAAAAUACAUCC-----------UAAUA-AAUUUCACAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUU--- ((((((((.....)))(((((((((((((...........-----------.....-.........))))....)))))))))..))))).......................--- ( -15.81) >DroSec_CAF1 50273 101 + 1 GUGGUGACUAAUUGUCGAUAAACAUGCUAAAAAUACAUGC-----------UACUA-AAUUUCAAAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUU--- ((((((((.....)))((((((((((.((.........((-----------((...-.........)))))).))))))))))..))))).......................--- ( -17.50) >DroSim_CAF1 53797 101 + 1 GUGGUGACUAAUUGUCGAUAAACAUGCUAAAAAUACAUGA-----------UACUA-AAUUUCAAAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUU--- ((((((((.....)))((((((((((.((....((..(((-----------.....-....)))..))..)).))))))))))..))))).......................--- ( -17.30) >DroEre_CAF1 52216 116 + 1 GUGGUCCAAAAUUGUCACUGAACAUGCUAAAAUCACAUGCCCGUGGUAAGCUAAUAACAUUUCGCAUAGCUACUGUGUUUAGUUUAGCACUUGAUUAAAGCAACAUCUACUGUUUA ((((........(((......)))((((..(((((.......(((.((((((((.........((((((...)))))))))))))).))).)))))..))))....))))...... ( -20.40) >consensus GUGGUGACUAAUUGUCGAUAAACAUGCUAAAAAUACAUGC___________UAAUA_AAUUUCAAAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUU___ (((((((.........(((((((((((((.....................................))))....)))))))))))))))).......................... (-12.30 = -12.12 + -0.19)



| Location | 10,518,797 – 10,518,893 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 97.92 |
| Mean single sequence MFE | -15.83 |
| Consensus MFE | -14.83 |
| Energy contribution | -15.17 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.683314 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10518797 96 + 22224390 UAAUAAAUUUCACAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUUUUAGCCUAACUAAAUGGUUAGGCAAUUGUGGAUUA ........((((((.........(((((...(((.....)))...)))))..............((((((((....))))))))...))))))... ( -18.20) >DroSec_CAF1 50313 96 + 1 UACUAAAUUUCAAAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUUUUAGCCUAACUAAAUGGUUAGGCAAUUGUGGAUUA .......................(((((...(((.....)))...)))))..((((((...((.((((((((....))))))))))..)))))).. ( -17.40) >DroSim_CAF1 53837 96 + 1 UACUAAAUUUCAAAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUUUUAGCCUAACUAAAUGGUUAGUCAAUUGUGGAUUA ..................................(((((((((((...........(((.(((((.....))))).))))))))))..)))).... ( -11.90) >consensus UACUAAAUUUCAAAUAGCUAACAUGUUUAUUUUACCACUUGAUUAAAACAACAUCUACUUUUUAGCCUAACUAAAUGGUUAGGCAAUUGUGGAUUA .......................(((((...(((.....)))...)))))..((((((...((.((((((((....))))))))))..)))))).. (-14.83 = -15.17 + 0.33)

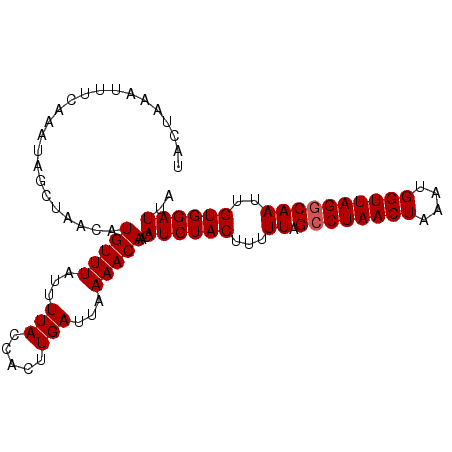

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:52 2006