
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,496,337 – 10,496,436 |
| Length | 99 |
| Max. P | 0.875664 |

| Location | 10,496,337 – 10,496,436 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 92.41 |
| Mean single sequence MFE | -35.16 |
| Consensus MFE | -32.19 |
| Energy contribution | -32.00 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.875664 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10496337 99 + 22224390 CCCACUCAUUUCGAAGCUCCGCUUUUCUGUGCUAAUUUCUGCUCCAGUUACGCGGCGGGCAAAUUGGCGAACAUUUUGCGGA-----UGGAGUGAAUGGAUGGG .(((.((((((((....(((((.......((((((((((((((.(......).))))))..))))))))........)))))-----)))))))).)))..... ( -32.86) >DroSec_CAF1 27483 104 + 1 CCCACUCAUUUCCAAGCUUCGCUUUUCUGUGCUAAUUUCUGCUCCAGUUACGCGGCUGGCAAAUUGGUGAGCAUUUUGCGGAUGGGAUGGAGUGAAUGGGUGGG (((((((((((....((((((((..(((((((((((((.....(((((......))))).)))))))).........)))))..)).))))))))))))))))) ( -36.70) >DroSim_CAF1 28009 104 + 1 CCCACUCAUUUCCAAGCUUCGCUUUUCUGUGCUAAUUUCUGCUCCAGUUACGCGGCGGGCAAAUUGGUGAGCAUUUUGCGGAUGGGAUGGAGUGAAUGGGUGGG (((((((((((....((((((((..((((((((((((((((((.(......).))))))..))))))).........)))))..)).))))))))))))))))) ( -37.00) >DroEre_CAF1 29071 99 + 1 CCCACUCAUUUUCAAGCUUCGCUUUUCUGUGCUAAUUUCUGCUCCAGUUACGCGGCGGGAAAAUUGGUGAGCAUUUUGCGAA-----UGGAGUGAAUGGGUGGG (((((((((((......(((((.....((((((((((((((((.(......).)))))))..))))))..)))....)))))-----......))))))))))) ( -34.10) >consensus CCCACUCAUUUCCAAGCUUCGCUUUUCUGUGCUAAUUUCUGCUCCAGUUACGCGGCGGGCAAAUUGGUGAGCAUUUUGCGGA_____UGGAGUGAAUGGGUGGG ((((((((((((((...(((((.....((((((((((((((((.(......).))))))..)))))))..)))....))))).....))))...)))))))))) (-32.19 = -32.00 + -0.19)


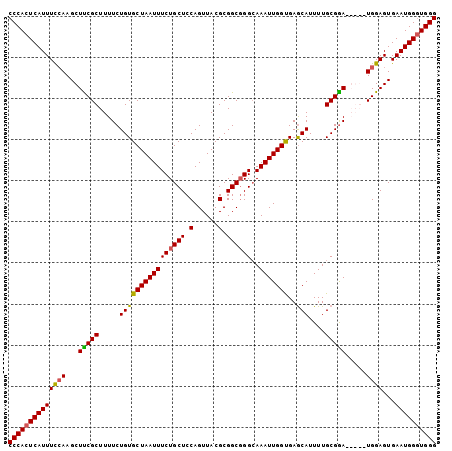
| Location | 10,496,337 – 10,496,436 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 92.41 |
| Mean single sequence MFE | -27.16 |
| Consensus MFE | -23.11 |
| Energy contribution | -23.68 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.678271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10496337 99 - 22224390 CCCAUCCAUUCACUCCA-----UCCGCAAAAUGUUCGCCAAUUUGCCCGCCGCGUAACUGGAGCAGAAAUUAGCACAGAAAAGCGGAGCUUCGAAAUGAGUGGG ((((..((((.......-----(((((....(((.(.....((((((((.........))).))))).....).))).....))))).......))))..)))) ( -25.74) >DroSec_CAF1 27483 104 - 1 CCCACCCAUUCACUCCAUCCCAUCCGCAAAAUGCUCACCAAUUUGCCAGCCGCGUAACUGGAGCAGAAAUUAGCACAGAAAAGCGAAGCUUGGAAAUGAGUGGG ....((((((((.((((........(((((.((.....)).))))).((((((....(((..((........)).)))....)))..)))))))..)))))))) ( -29.20) >DroSim_CAF1 28009 104 - 1 CCCACCCAUUCACUCCAUCCCAUCCGCAAAAUGCUCACCAAUUUGCCCGCCGCGUAACUGGAGCAGAAAUUAGCACAGAAAAGCGAAGCUUGGAAAUGAGUGGG ....((((((((.((((........(((((.((.....)).)))))..(((((....(((..((........)).)))....)))..)).))))..)))))))) ( -28.70) >DroEre_CAF1 29071 99 - 1 CCCACCCAUUCACUCCA-----UUCGCAAAAUGCUCACCAAUUUUCCCGCCGCGUAACUGGAGCAGAAAUUAGCACAGAAAAGCGAAGCUUGAAAAUGAGUGGG ....(((((((......-----...((.....))......((((((..(((((....(((..((........)).)))....)))..))..))))))))))))) ( -25.00) >consensus CCCACCCAUUCACUCCA_____UCCGCAAAAUGCUCACCAAUUUGCCCGCCGCGUAACUGGAGCAGAAAUUAGCACAGAAAAGCGAAGCUUGGAAAUGAGUGGG ....((((((((.((((........(((((.((.....)).)))))..(((((....(((..((........)).)))....)))..)).))))..)))))))) (-23.11 = -23.68 + 0.56)

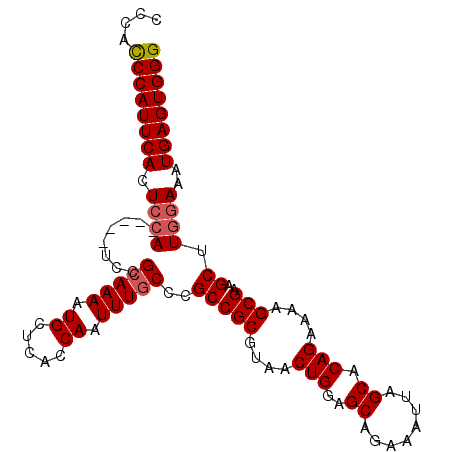

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:21 2006