
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,453,979 – 10,454,125 |
| Length | 146 |
| Max. P | 0.992588 |

| Location | 10,453,979 – 10,454,098 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -27.05 |
| Consensus MFE | -25.66 |
| Energy contribution | -25.85 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.95 |
| SVM decision value | 2.34 |
| SVM RNA-class probability | 0.992588 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10453979 119 + 22224390 GGCGCUGCAGUGCCGCUCGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAU-UUCGGAUUGU (((((....)))))....(((((((..((((((((........(((((...(.....)..))))).(((((..((..((......))...))..))))).....)))))-)))))))))) ( -28.90) >DroVir_CAF1 14875 120 + 1 GGCGCUGCAAUGCCGCACGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAUUUUCGGAUUGU ((((......))))....(((((((...(((((........))))).....((((((.((((....(((((..((..((......))...))..)))))...)))).))))))))))))) ( -25.70) >DroGri_CAF1 12691 120 + 1 GGCGCUGCAAUGCCGCACGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAUUUUCGGAUUGU ((((......))))....(((((((...(((((........))))).....((((((.((((....(((((..((..((......))...))..)))))...)))).))))))))))))) ( -25.70) >DroWil_CAF1 11772 120 + 1 GGCGCUGCAAUGCCGCACGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAUUUUCGGAUUGU ((((......))))....(((((((...(((((........))))).....((((((.((((....(((((..((..((......))...))..)))))...)))).))))))))))))) ( -25.70) >DroMoj_CAF1 13275 120 + 1 GGCGCUGCAGUGCCGCACGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAUUUUCGGAUUGC (((((....)))))....(((((((...(((((........))))).....((((((.((((....(((((..((..((......))...))..)))))...)))).))))))))))))) ( -31.00) >DroAna_CAF1 16385 119 + 1 GGCGUUGAAGUGCCGCUCGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAU-UUCGGAUUGU ((((......))))....(((((((..((((((((........(((((...(.....)..))))).(((((..((..((......))...))..))))).....)))))-)))))))))) ( -25.30) >consensus GGCGCUGCAAUGCCGCACGCGAUCCUAGAAGUUUUGAACAUAAUUUACACUGAAAAUCUCGUAAAACUUCAGCCAUAUCUAAAAUGAAAUUGUAUGAAGAAAAUGAAAUUUUCGGAUUGU ((((......))))....(((((((...(((((........))))).....((((((.((((....(((((..((..((......))...))..)))))...)))).))))))))))))) (-25.66 = -25.85 + 0.19)

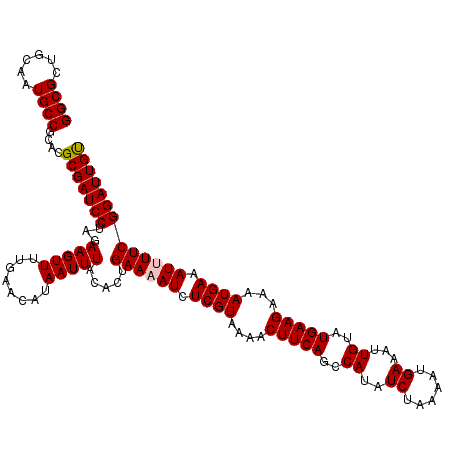

| Location | 10,454,019 – 10,454,125 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.53 |
| Mean single sequence MFE | -15.08 |
| Consensus MFE | -13.85 |
| Energy contribution | -14.05 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.619975 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10454019 106 - 22224390 ---UCACUAAGUU----GUUCUUU------CAAAAUCACCACAAUCCGAA-AUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA ---.((((..(((----((.....------..........)))))..(((-(((((.....(((((..((...((......))..))..)))))......)))))))).))))....... ( -12.96) >DroPse_CAF1 13740 114 - 1 AAUUCACUAAGUUUCUUCUUCUUU------CAAAAUCACCACAAUCCGAAAAUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA ....(((((((......)))....------.................(((((((((.....(((((..((...((......))..))..)))))......)))))))))))))....... ( -16.30) >DroWil_CAF1 11812 110 - 1 --AUCACUAAUUU----CCUUUUU---CU-UAAAAUCACCACAAUCCGAAAAUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA --..((((.....----..((((.---..-.))))............(((((((((.....(((((..((...((......))..))..)))))......)))))))))))))....... ( -15.50) >DroMoj_CAF1 13315 106 - 1 --CUCACUAAUUU------UCUUU------CAAAAUCACCGCAAUCCGAAAAUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA --..((((.((((------(....------.)))))...........(((((((((.....(((((..((...((......))..))..)))))......)))))))))))))....... ( -15.60) >DroAna_CAF1 16425 112 - 1 ---UCACUAAGUU----GUUACUUUUUAUAAAAAAUCACCACAAUCCGAA-AUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA ---.((((..(((----((....((((....)))).....)))))..(((-(((((.....(((((..((...((......))..))..)))))......)))))))).))))....... ( -13.80) >DroPer_CAF1 40452 114 - 1 AAUUCACUAAGUUUCUUCUUCUUU------CAAAAUCACCACAAUCCGAAAAUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA ....(((((((......)))....------.................(((((((((.....(((((..((...((......))..))..)))))......)))))))))))))....... ( -16.30) >consensus ___UCACUAAGUU____CUUCUUU______CAAAAUCACCACAAUCCGAAAAUUUCAUUUUCUUCAUACAAUUUCAUUUUAGAUAUGGCUGAAGUUUUACGAGAUUUUCAGUGUAAAUUA ........................................(((....(((((((((.....(((((..((...((......))..))..)))))......)))))))))..)))...... (-13.85 = -14.05 + 0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:11:21 2006