
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,393,152 – 10,393,249 |
| Length | 97 |
| Max. P | 0.762274 |

| Location | 10,393,152 – 10,393,249 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 79.19 |
| Mean single sequence MFE | -27.01 |
| Consensus MFE | -16.21 |
| Energy contribution | -16.57 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.762274 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
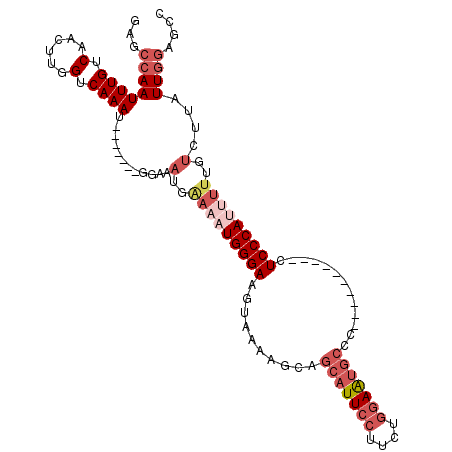
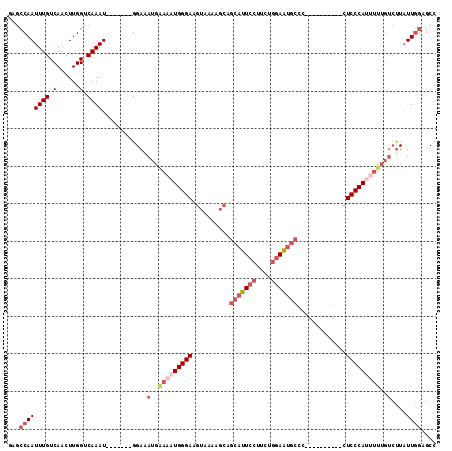
>X_DroMel_CAF1 10393152 97 + 22224390 GGCUCCAAUAAGACAAAUAUGGGAG----------GGGCAUUCCAGAAGGAAUGCUGCUUUUACUUCCCAUUUCCAUUUCC-------AUUUGACCAAGCUGACAAAUUGGCUG ((.((.(((.........(((((((----------(((((((((....)))))))).......))))))))..........-------))).)))).(((..(....)..))). ( -27.32) >DroSec_CAF1 37147 97 + 1 GGCUCCAAUAAGACAAAACUGGGAG----------GGGCAUUCCAGAAGGAAUGCUGCUUUUACUUCCCAUUUCCAUUUCC-------AUUUGACCAAGCUGACAAAUUGGCUA ((.((.(((..........((((((----------(((((((((....)))))))).......)))))))...........-------))).)))).(((..(....)..))). ( -25.41) >DroEre_CAF1 41783 97 + 1 GGCUCCAAUAAGACAAAAAUGGGAG----------AGGCAUUCCAGAAGGAAUGCUGCUUUUACUUCCCAUUUUCAUUUCC-------AUUUGACCAAGUUGACAAAUUGGCUC ((.((.(((.(((..((((((((((----------.((((((((....))))))))........))))))))))..)))..-------))).)))).(((..(....)..))). ( -26.20) >DroYak_CAF1 36865 97 + 1 GGCUCCAAUAAGACAAAAAUGGGAG----------GGGCAUUCCCGAAGGAAUGCUGCUUUUACUUCCCAUUUUCAUUUCC-------AUUUGACCAAGUUGACAAAUUGGCUC ((.((.(((.(((..((((((((((----------(((((((((....)))))))).......)))))))))))..)))..-------))).)))).(((..(....)..))). ( -28.41) >DroAna_CAF1 31632 106 + 1 GGCUUUAAGAG----AGUGUGGGAGAGAAAGAGUAAGGUGCUUGAGAAGGAAUGUGACUUC----UCCCAUUUACUUUUCCUUCUGCUAUUUGGCCAAGUUGACAAAUUGUUUC ((((......(----((((..((((.(((.(((((((.((...((((((........))))----)).))))))))))))))))..).)))))))).................. ( -27.70) >consensus GGCUCCAAUAAGACAAAAAUGGGAG__________GGGCAUUCCAGAAGGAAUGCUGCUUUUACUUCCCAUUUCCAUUUCC_______AUUUGACCAAGUUGACAAAUUGGCUC ................(((((((((...........((((((((....))))))))........)))))))))........................((((((....)))))). (-16.21 = -16.57 + 0.36)



| Location | 10,393,152 – 10,393,249 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 79.19 |
| Mean single sequence MFE | -25.54 |
| Consensus MFE | -16.50 |
| Energy contribution | -18.98 |
| Covariance contribution | 2.48 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.587782 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10393152 97 - 22224390 CAGCCAAUUUGUCAGCUUGGUCAAAU-------GGAAAUGGAAAUGGGAAGUAAAAGCAGCAUUCCUUCUGGAAUGCCC----------CUCCCAUAUUUGUCUUAUUGGAGCC ...(((((..(.(((((...((....-------.))...))..((((((.((....)).(((((((....)))))))..----------.))))))..))).)..))))).... ( -25.80) >DroSec_CAF1 37147 97 - 1 UAGCCAAUUUGUCAGCUUGGUCAAAU-------GGAAAUGGAAAUGGGAAGUAAAAGCAGCAUUCCUUCUGGAAUGCCC----------CUCCCAGUUUUGUCUUAUUGGAGCC ...(((((..(.(((((...((....-------.))...))...(((((.((....)).(((((((....)))))))..----------.)))))...))).)..))))).... ( -24.80) >DroEre_CAF1 41783 97 - 1 GAGCCAAUUUGUCAACUUGGUCAAAU-------GGAAAUGAAAAUGGGAAGUAAAAGCAGCAUUCCUUCUGGAAUGCCU----------CUCCCAUUUUUGUCUUAUUGGAGCC (..((((((((.(......).)))))-------.((.(..(((((((((.((....)).(((((((....)))))))..----------.)))))))))..).))..)))..). ( -28.90) >DroYak_CAF1 36865 97 - 1 GAGCCAAUUUGUCAACUUGGUCAAAU-------GGAAAUGAAAAUGGGAAGUAAAAGCAGCAUUCCUUCGGGAAUGCCC----------CUCCCAUUUUUGUCUUAUUGGAGCC (..((((((((.(......).)))))-------.((.(..(((((((((.((....)).(((((((....)))))))..----------.)))))))))..).))..)))..). ( -31.70) >DroAna_CAF1 31632 106 - 1 GAAACAAUUUGUCAACUUGGCCAAAUAGCAGAAGGAAAAGUAAAUGGGA----GAAGUCACAUUCCUUCUCAAGCACCUUACUCUUUCUCUCCCACACU----CUCUUAAAGCC ......(((((((.....)).)))))....((..(((((((((.((.((----((((........))))))...))..))))).))))..)).......----........... ( -16.50) >consensus GAGCCAAUUUGUCAACUUGGUCAAAU_______GGAAAUGAAAAUGGGAAGUAAAAGCAGCAUUCCUUCUGGAAUGCCC__________CUCCCAUUUUUGUCUUAUUGGAGCC ...((((((((.(......).))))............(..(((((((((..........(((((((....))))))).............)))))))))..)....)))).... (-16.50 = -18.98 + 2.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:11:06 2006