
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,211,851 – 10,211,971 |
| Length | 120 |
| Max. P | 0.857719 |

| Location | 10,211,851 – 10,211,971 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.42 |
| Mean single sequence MFE | -43.02 |
| Consensus MFE | -37.04 |
| Energy contribution | -37.32 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597148 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10211851 120 + 22224390 ACCACAUUGGUGACCGCGUAGUUGCAAGCCAGCAGGGUGGCCUGCACAUUGUUGGCGUCCGUAAAUCGAGCAAUCGCACAACCCACUCGGUUAUUUCGCUCGUUGACCAUGGUGGUGAAG (((((..(((((((.(((..(((((..((((((((.(((.....))).))))))))...((.....)).)))))..)..((((.....)))).....))..))).))))..))))).... ( -41.40) >DroSec_CAF1 8180 120 + 1 ACCACAUUGGUGACCGCGUAGUUGCAGGCCAACAGGGUGGCCUGCACAUUGUUGGCGUCCGUAAAUCGGGCAAUCGCACAUCCAACUCGGUUAUUUCGCUCGUUGACCAUGGUGGUGAAG (((((..((((....((((((((((((((((......))))))))).)))).....(((((.....)))))...))).....((((.(((.....)))...))))))))..))))).... ( -43.70) >DroSim_CAF1 8649 120 + 1 ACCACAUUGGUGACCGCGUAGUUGCAGGCCAACAGGGUGGCCUGCACAUUGUUGGCGUCCGUAAAUCGGGCAAUCGCACAUCCCACUCGGUUAUUUCGCUCGUUGACCAUGGUGGUGAAG (((.....))).(((((.....(((((((((......)))))))))(((.((..(((((((.....)))))(((((...........))))).........))..)).)))))))).... ( -43.00) >DroEre_CAF1 9343 120 + 1 AGCACAUUGGUGACCGCGUAAUUGCAGGCCAACAGGGUGGCCUGCACGUUGUUGGCGUCCGUAAAUCGGGCAAUCGCACAACCCACUCUGUUGUUACGCUCGUUGACCAUGGUGGUGAAG ..(((..(((((((.(((((((.((((((((((((.(((.....))).))))))))(((((.....)))))................)))).)))))))..))).))))..)))...... ( -48.80) >DroYak_CAF1 9327 120 + 1 AACACAUUGGUGACCGCGUAAUUGCAGACCAACAGAGUGGCCUGCACGUUAUUGGCGUCCGUAAAACGGGCAAUCGCACAUCCCACUCGGUUAUUACGUUCGUUGACCAUCGUGGUGAAG ..(((..(((((((.(((((((....((((....((((((..(((..((.....))(((((.....)))))....)))....)))))))))))))))))..))).))))..)))...... ( -38.20) >consensus ACCACAUUGGUGACCGCGUAGUUGCAGGCCAACAGGGUGGCCUGCACAUUGUUGGCGUCCGUAAAUCGGGCAAUCGCACAUCCCACUCGGUUAUUUCGCUCGUUGACCAUGGUGGUGAAG (((((..(((((((.(((.(((.((..((((((((.(((.....))).))))))))(((((.....)))))..................)).))).)))..))).))))..))))).... (-37.04 = -37.32 + 0.28)


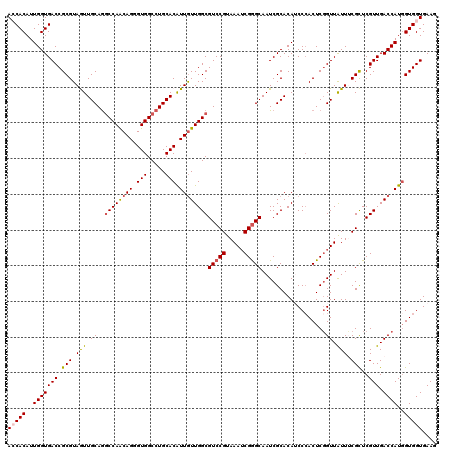
| Location | 10,211,851 – 10,211,971 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.42 |
| Mean single sequence MFE | -41.58 |
| Consensus MFE | -35.58 |
| Energy contribution | -36.10 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.857719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10211851 120 - 22224390 CUUCACCACCAUGGUCAACGAGCGAAAUAACCGAGUGGGUUGUGCGAUUGCUCGAUUUACGGACGCCAACAAUGUGCAGGCCACCCUGCUGGCUUGCAACUACGCGGUCACCAAUGUGGU ....(((((..((((...(((((((.((((((.....))))))....)))))))......(..(((........(((((((((......))))))))).....)))..)))))..))))) ( -44.92) >DroSec_CAF1 8180 120 - 1 CUUCACCACCAUGGUCAACGAGCGAAAUAACCGAGUUGGAUGUGCGAUUGCCCGAUUUACGGACGCCAACAAUGUGCAGGCCACCCUGUUGGCCUGCAACUACGCGGUCACCAAUGUGGU ....(((((..((((..((..(((..........(((((.(((((....))(((.....)))))))))))....(((((((((......)))))))))....))).)).))))..))))) ( -43.10) >DroSim_CAF1 8649 120 - 1 CUUCACCACCAUGGUCAACGAGCGAAAUAACCGAGUGGGAUGUGCGAUUGCCCGAUUUACGGACGCCAACAAUGUGCAGGCCACCCUGUUGGCCUGCAACUACGCGGUCACCAAUGUGGU ....(((((..((((......(((......((....))....)))(((((((((.....)))............(((((((((......))))))))).....))))))))))..))))) ( -40.60) >DroEre_CAF1 9343 120 - 1 CUUCACCACCAUGGUCAACGAGCGUAACAACAGAGUGGGUUGUGCGAUUGCCCGAUUUACGGACGCCAACAACGUGCAGGCCACCCUGUUGGCCUGCAAUUACGCGGUCACCAAUGUGCU ......(((..((((..((..((((((.......((..((((.(((.....(((.....))).))))))).)).(((((((((......))))))))).)))))).)).))))..))).. ( -41.20) >DroYak_CAF1 9327 120 - 1 CUUCACCACGAUGGUCAACGAACGUAAUAACCGAGUGGGAUGUGCGAUUGCCCGUUUUACGGACGCCAAUAACGUGCAGGCCACUCUGUUGGUCUGCAAUUACGCGGUCACCAAUGUGUU ......((((.((((............((((.((((((..((..((((((((((.....)))..).))))..))..))..)))))).))))(.((((......)))).))))).)))).. ( -38.10) >consensus CUUCACCACCAUGGUCAACGAGCGAAAUAACCGAGUGGGAUGUGCGAUUGCCCGAUUUACGGACGCCAACAAUGUGCAGGCCACCCUGUUGGCCUGCAACUACGCGGUCACCAAUGUGGU ....(((((..((((...................((((((((.((....)).))))))))(..(((........(((((((((......))))))))).....)))..)))))..))))) (-35.58 = -36.10 + 0.52)

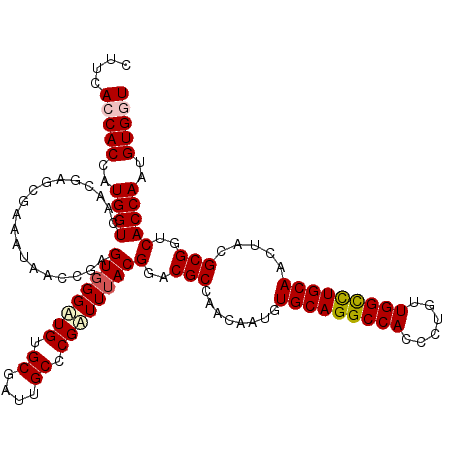
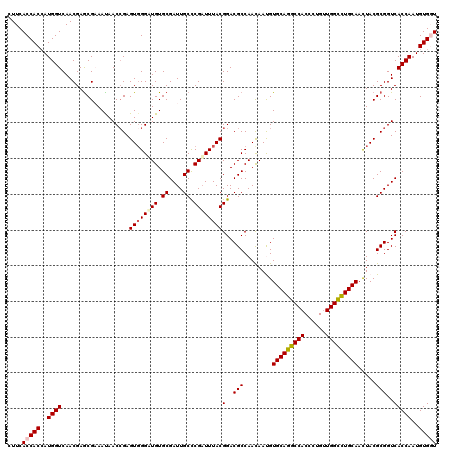
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:10:04 2006