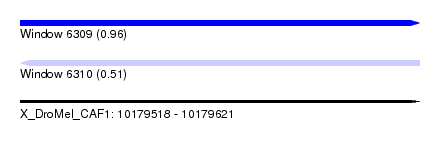

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,179,518 – 10,179,621 |
| Length | 103 |
| Max. P | 0.958453 |

| Location | 10,179,518 – 10,179,621 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 78.92 |
| Mean single sequence MFE | -22.00 |
| Consensus MFE | -16.70 |
| Energy contribution | -17.70 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.49 |
| SVM RNA-class probability | 0.958453 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10179518 103 + 22224390 ------AUUAUAAUAAUGAUA-------ACAACUACGUGAUAAUCUUCCAUAGAAGACUCCACCGGACUCUUUUAAGCUGUAAAAAGGAAGUAGACUAGUAGACUAUCUGGUUGUU ------..............(-------((((((..........(((((.(((((((.(((...))).)))))))...(......)))))).(((.(((....))))))))))))) ( -18.40) >DroSec_CAF1 8147 100 + 1 AAUUUAAUUAUAAUAAUGAUA-------ACAACUACGUGAUAAUCUUCCAUAGAAGACUCCGCAAGAGUCUUUUAAGCUAUUAAAAGCAUGUAG---------UUAUCUGGUUGUU .........(((((...((((-------...(((((((............(((((((((.(....)))))))))).(((......)))))))))---------)))))..))))). ( -24.60) >DroSim_CAF1 9289 107 + 1 AAUUUAAUUAUAAUAAUGACUACGUGAUACAACUACCUGAUAAUCUUCCAUAGAAGACUCCGCAAGAGUCUUUUAAGCUAUUAAAAGCAAGUAG---------UUUUCUGGUUGUU ..............((..((((...((...((((((..............(((((((((.(....)))))))))).(((......)))..))))---------)).))))))..)) ( -23.00) >consensus AAUUUAAUUAUAAUAAUGAUA_______ACAACUACGUGAUAAUCUUCCAUAGAAGACUCCGCAAGAGUCUUUUAAGCUAUUAAAAGCAAGUAG_________UUAUCUGGUUGUU ............................(((((((...((((........((((((((((.....)))))))))).(((......)))................))))))))))). (-16.70 = -17.70 + 1.00)

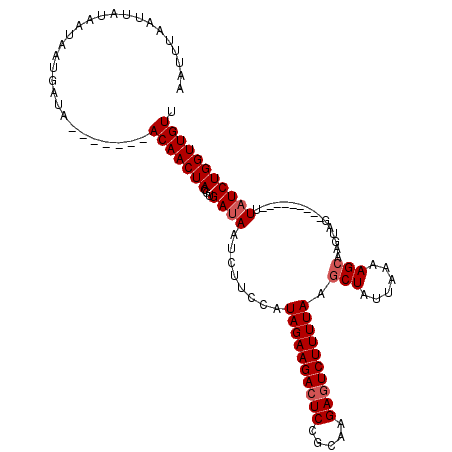
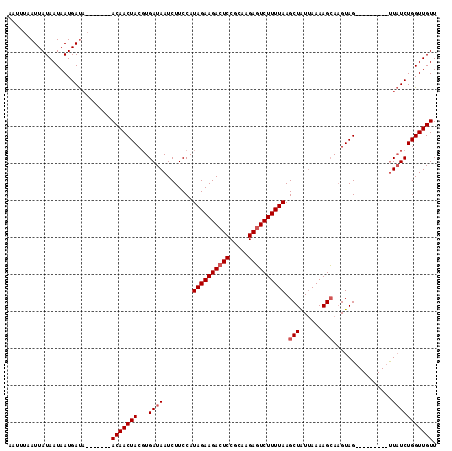
| Location | 10,179,518 – 10,179,621 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 78.92 |
| Mean single sequence MFE | -21.10 |
| Consensus MFE | -13.13 |
| Energy contribution | -14.13 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.62 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.512586 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10179518 103 - 22224390 AACAACCAGAUAGUCUACUAGUCUACUUCCUUUUUACAGCUUAAAAGAGUCCGGUGGAGUCUUCUAUGGAAGAUUAUCACGUAGUUGU-------UAUCAUUAUUAUAAU------ ........(((((.((((...((((((..((((((........))))))...))))))((((((....))))))......))))...)-------))))...........------ ( -20.40) >DroSec_CAF1 8147 100 - 1 AACAACCAGAUAA---------CUACAUGCUUUUAAUAGCUUAAAAGACUCUUGCGGAGUCUUCUAUGGAAGAUUAUCACGUAGUUGU-------UAUCAUUAUUAUAAUUAAAUU ........(((((---------(.((...(((((((....))))))).....((((.(((((((....)))))))....)))))).))-------))))................. ( -21.80) >DroSim_CAF1 9289 107 - 1 AACAACCAGAAAA---------CUACUUGCUUUUAAUAGCUUAAAAGACUCUUGCGGAGUCUUCUAUGGAAGAUUAUCAGGUAGUUGUAUCACGUAGUCAUUAUUAUAAUUAAAUU ........((.((---------(((((((((((((.(((.....(((((((.....)))))))))))))))).....)))))))))...))......................... ( -21.10) >consensus AACAACCAGAUAA_________CUACUUGCUUUUAAUAGCUUAAAAGACUCUUGCGGAGUCUUCUAUGGAAGAUUAUCACGUAGUUGU_______UAUCAUUAUUAUAAUUAAAUU ........((((..........((((...(((((((....)))))))........(((((((((....))))))).))..))))...........))))................. (-13.13 = -14.13 + 1.00)

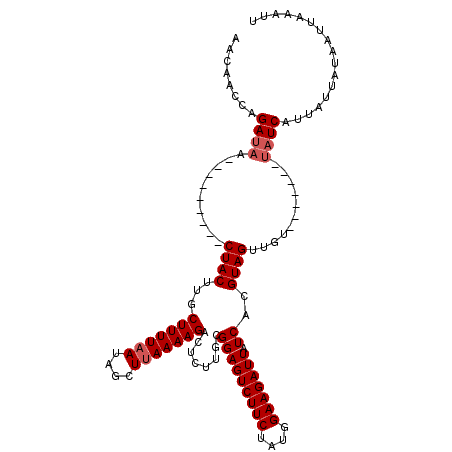
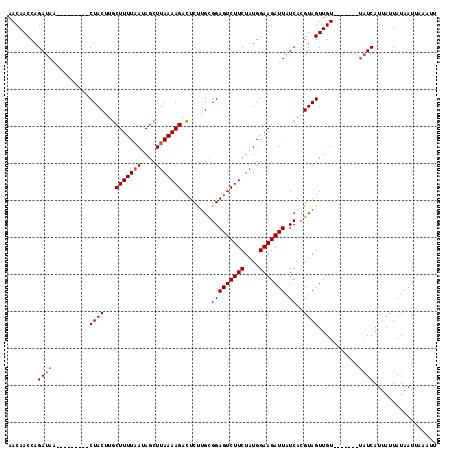
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:09:53 2006