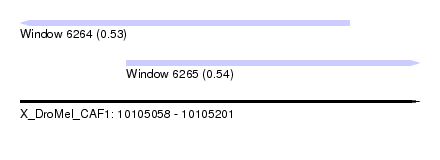
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,105,058 – 10,105,201 |
| Length | 143 |
| Max. P | 0.537281 |

| Location | 10,105,058 – 10,105,176 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.58 |
| Mean single sequence MFE | -35.80 |
| Consensus MFE | -31.18 |
| Energy contribution | -32.73 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.87 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532557 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10105058 118 - 22224390 GCAGCCAGCCACUUAACCCUCGAGUGGCUCUCGCUGAUGUAAUGCCAUUCGAUUUGAUUUGCCAUGGUUAACCAGAUUUGCUUGCCACAGCCAUUCGGCAUUUUGCGGUUGCUAAUGC-- ((((((.((..........(((((((((....((....))...)))))))))...((..((((.(((....)))((...(((......)))...))))))..))))))))))......-- ( -37.80) >DroYak_CAF1 21371 118 - 1 GCAGCCAGCCACUUAACCCUCGAGUGGCUCUCGCUGAUGUAAUGCCAUUCGAUUUGAUUUGCCAUGGUUAACCAGAUUUGCUUGCCACAGCCAUUCGGCAUUUUGCAGCUGCUAUUGC-- (((((..((..........(((((((((....((....))...)))))))))...((..((((.(((....)))((...(((......)))...))))))..)))).)))))......-- ( -36.20) >DroAna_CAF1 35478 117 - 1 G---GCAGCCACUUAACCCUCGAGUGGCUCUCGCUGAUGUAAUGCCAUUCGAUUUGAUUUGCCAUGGUUAACCAGAUUUGCUUGCCACAGCCAUUCGGCAUUUUGCGGUUCCUAGUCCUU (---(.((((.........(((((((((....((....))...)))))))))...((..((((.(((....)))((...(((......)))...))))))..))..))))))........ ( -33.40) >consensus GCAGCCAGCCACUUAACCCUCGAGUGGCUCUCGCUGAUGUAAUGCCAUUCGAUUUGAUUUGCCAUGGUUAACCAGAUUUGCUUGCCACAGCCAUUCGGCAUUUUGCGGUUGCUAAUGC__ ((((((.((..........(((((((((....((....))...)))))))))...((..((((.(((....)))((...(((......)))...))))))..))))))))))........ (-31.18 = -32.73 + 1.56)


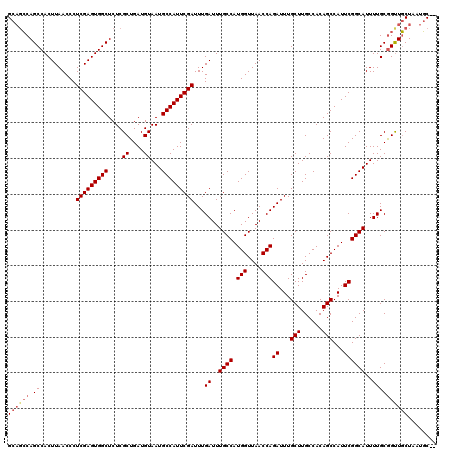
| Location | 10,105,096 – 10,105,201 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.67 |
| Mean single sequence MFE | -31.61 |
| Consensus MFE | -25.57 |
| Energy contribution | -26.23 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537281 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
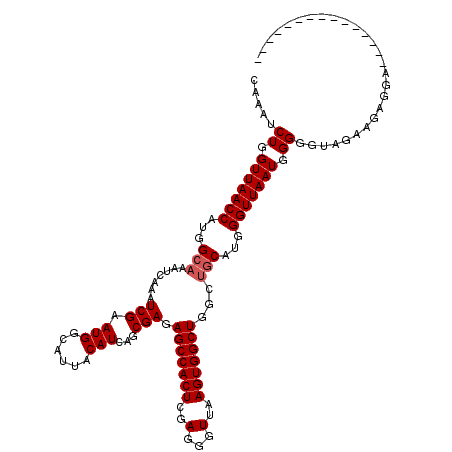
>X_DroMel_CAF1 10105096 105 + 22224390 CAAAUCUGGUUAACCAUGGCAAAUCAAAUCGAAUGGCAUUACAUCAGCGAGAGCCACUCGAGGGUUAAGUGGCUGGCUGCAUGGGUUAAUGGGGGUAGAAGAGGA--------------- ....(((.(((((((...((...((.....))...))....(((((((...(((((((..(...)..))))))).)))).)))))))))).)))...........--------------- ( -30.80) >DroYak_CAF1 21409 105 + 1 CAAAUCUGGUUAACCAUGGCAAAUCAAAUCGAAUGGCAUUACAUCAGCGAGAGCCACUCGAGGGUUAAGUGGCUGGCUGCAUGGGUUAAUGGGGGUAGAAGAGGA--------------- ....(((.(((((((...((...((.....))...))....(((((((...(((((((..(...)..))))))).)))).)))))))))).)))...........--------------- ( -30.80) >DroAna_CAF1 35518 116 + 1 CAAAUCUGGUUAACCAUGGCAAAUCAAAUCGAAUGGCAUUACAUCAGCGAGAGCCACUCGAGGGUUAAGUGGCUGC---CAUGGGUUAAUGGG-CAACAAGAGGAACACGACCCACCAUA ......((((...(((((((........(((.(((......)))...))).(((((((..(...)..)))))))))---)))))......(((-(..............).))))))).. ( -33.24) >consensus CAAAUCUGGUUAACCAUGGCAAAUCAAAUCGAAUGGCAUUACAUCAGCGAGAGCCACUCGAGGGUUAAGUGGCUGGCUGCAUGGGUUAAUGGGGGUAGAAGAGGA_______________ .....((.(((((((...(((.......(((.(((......)))...))).(((((((..(...)..)))))))...)))...))))))).))........................... (-25.57 = -26.23 + 0.67)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:09:14 2006