| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,029,100 – 10,029,203 |
| Length | 103 |
| Max. P | 0.547641 |

| Location | 10,029,100 – 10,029,203 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.84 |
| Mean single sequence MFE | -24.50 |
| Consensus MFE | -16.99 |
| Energy contribution | -16.77 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516462 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10029100 103 + 22224390 AUGCAAAAAGUUGUAAGCGAAAAGCCAGAGGUGCAAAGGAGGACCAUUGCAAAAAUCAACAGGCCAUAAUGGGUUAAUUAAGGCAGGG-----UGA----UGGCAAACUGAA-------- .(((((....))))).((.....))(((.(((.(......).))).((((....((((.(..(((.((((......)))).)))..).-----)))----).)))).)))..-------- ( -25.20) >DroEre_CAF1 36459 103 + 1 AUGCAAAAAGUUGUGCGCGAAAGACCAGAGGUGCAAAGGAGGACCAUUGCAAAAAUCAACAGGCCAUAAUGGGUUAAUUAAGGCAGGG-----UGA----UGGCAAACUGAA-------- ..(((........))).(....)..(((.(((.(......).))).((((....((((.(..(((.((((......)))).)))..).-----)))----).)))).)))..-------- ( -25.20) >DroYak_CAF1 33747 120 + 1 AUGCAAAAUGUUGUGAGCGAAGGGCGAGAGGUGCAAAGGAGGAGCAUUGCAAAAAUCAACAGGCCAUAAUGGGUUAAUUAAGGCAGUGAUGGCUGAUGGGUGGCAAACUGAAAACUGAGA .((((...(.((((...(....))))).)..))))...........((((....(((....((((((.....(((......)))....))))))....))).)))).............. ( -23.10) >consensus AUGCAAAAAGUUGUGAGCGAAAGGCCAGAGGUGCAAAGGAGGACCAUUGCAAAAAUCAACAGGCCAUAAUGGGUUAAUUAAGGCAGGG_____UGA____UGGCAAACUGAA________ .(((.....((((..........((.....))((((.(......).))))......))))..(((.((((......)))).)))..................)))............... (-16.99 = -16.77 + -0.22)


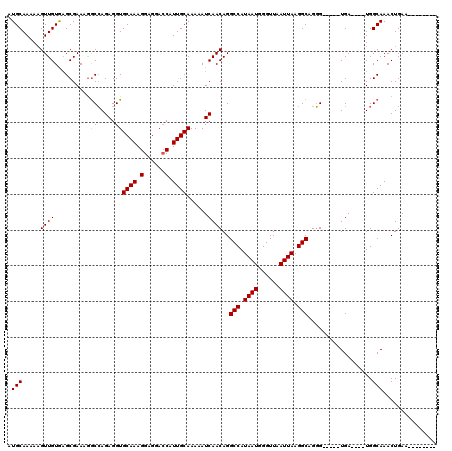
| Location | 10,029,100 – 10,029,203 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.84 |
| Mean single sequence MFE | -17.23 |
| Consensus MFE | -14.82 |
| Energy contribution | -14.93 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.547641 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10029100 103 - 22224390 --------UUCAGUUUGCCA----UCA-----CCCUGCCUUAAUUAACCCAUUAUGGCCUGUUGAUUUUUGCAAUGGUCCUCCUUUGCACCUCUGGCUUUUCGCUUACAACUUUUUGCAU --------........((((----(((-----.(..(((.((((......)))).)))..).)))....(((((.((....)).)))))....)))).....((............)).. ( -19.10) >DroEre_CAF1 36459 103 - 1 --------UUCAGUUUGCCA----UCA-----CCCUGCCUUAAUUAACCCAUUAUGGCCUGUUGAUUUUUGCAAUGGUCCUCCUUUGCACCUCUGGUCUUUCGCGCACAACUUUUUGCAU --------........((((----(((-----.(..(((.((((......)))).)))..).)))....(((((.((....)).)))))....)))).......(((........))).. ( -18.20) >DroYak_CAF1 33747 120 - 1 UCUCAGUUUUCAGUUUGCCACCCAUCAGCCAUCACUGCCUUAAUUAACCCAUUAUGGCCUGUUGAUUUUUGCAAUGCUCCUCCUUUGCACCUCUCGCCCUUCGCUCACAACAUUUUGCAU ...............(((..................(((.((((......)))).))).(((((.....(((((..........)))))......((.....))...)))))....))). ( -14.40) >consensus ________UUCAGUUUGCCA____UCA_____CCCUGCCUUAAUUAACCCAUUAUGGCCUGUUGAUUUUUGCAAUGGUCCUCCUUUGCACCUCUGGCCUUUCGCUCACAACUUUUUGCAU ...............(((..................(((.((((......)))).)))..((((.....(((((.((....)).)))))......((.....))...)))).....))). (-14.82 = -14.93 + 0.11)


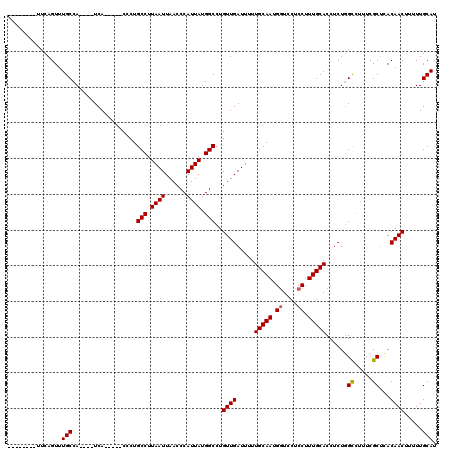
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:08:33 2006