
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,027,336 – 10,027,456 |
| Length | 120 |
| Max. P | 0.994811 |

| Location | 10,027,336 – 10,027,456 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.78 |
| Mean single sequence MFE | -23.23 |
| Consensus MFE | -17.48 |
| Energy contribution | -17.59 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.926126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10027336 120 + 22224390 AGCAUAAAUGGUAUUGAGUUAAUGAAUAACUCAAUUUUGGCCUAUUGUUACCCACAACAUAUUGUGCAACAUAUUACUCAUACGCACUGUUGCACCAUAUUGUAUUUAGCGACAACAAAU .((.(((((((((((((((((.....)))))))))....)))..............((((((.((((((((................)))))))).))).)))))))))).......... ( -25.39) >DroSim_CAF1 4656 120 + 1 AGCAUAUAUGAUAUUGAGUUAAUGAAUAACUCAAUUUUGGCUUAUUGCUAUCCACAACAUACUGUGCAACAUAUUACUCAUACGCCCUGUUGCACCAUAUUGUAUUCAGCGACAACAAAU .((..(((..(((((((((((.....))))))))...((((.....)))).............((((((((................))))))))..)))..)))...)).......... ( -24.99) >DroYak_CAF1 31866 107 + 1 -------------UUGAGUUAAUAAAUAACUCAAUUUUGCCUCAUUGUUAUCCGAAACAUGCAUUGCAACAUAUUACGCAUACGCCCUGUUGCACCAAAUUGUAUACAGCGACAACAAAU -------------((((((((.....))))))))..........(((((...((....(((((.(((((((......((....))..)))))))......)))))....))..))))).. ( -19.30) >consensus AGCAUA_AUG_UAUUGAGUUAAUGAAUAACUCAAUUUUGGCUUAUUGUUAUCCACAACAUACUGUGCAACAUAUUACUCAUACGCCCUGUUGCACCAUAUUGUAUUCAGCGACAACAAAU .............((((((((.....)))))))).((((.....(((((.......(((....((((((((................)))))))).....)))....)))))...)))). (-17.48 = -17.59 + 0.11)

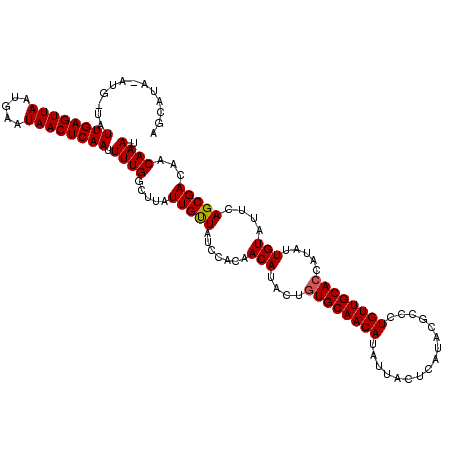
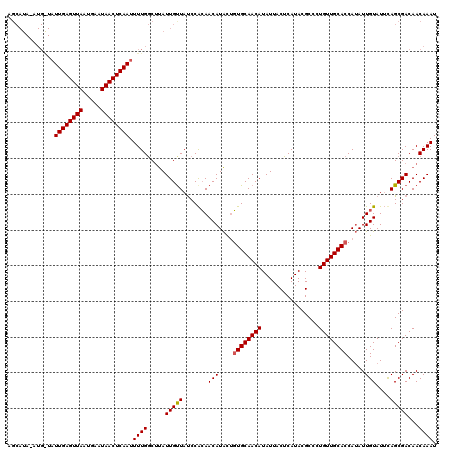
| Location | 10,027,336 – 10,027,456 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.78 |
| Mean single sequence MFE | -34.93 |
| Consensus MFE | -29.46 |
| Energy contribution | -29.13 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.17 |
| Mean z-score | -3.90 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.51 |
| SVM RNA-class probability | 0.994811 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10027336 120 - 22224390 AUUUGUUGUCGCUAAAUACAAUAUGGUGCAACAGUGCGUAUGAGUAAUAUGUUGCACAAUAUGUUGUGGGUAACAAUAGGCCAAAAUUGAGUUAUUCAUUAACUCAAUACCAUUUAUGCU .((((((((..((...(((((((((((((((((.(((......)))...))))))))..)))))))))))..)))))))).....(((((((((.....)))))))))............ ( -38.20) >DroSim_CAF1 4656 120 - 1 AUUUGUUGUCGCUGAAUACAAUAUGGUGCAACAGGGCGUAUGAGUAAUAUGUUGCACAGUAUGUUGUGGAUAGCAAUAAGCCAAAAUUGAGUUAUUCAUUAACUCAAUAUCAUAUAUGCU .((((((((.((((..(((((((((((((((((..((......))....))))))))..)))))))))..)))).))))..))))(((((((((.....)))))))))............ ( -36.90) >DroYak_CAF1 31866 107 - 1 AUUUGUUGUCGCUGUAUACAAUUUGGUGCAACAGGGCGUAUGCGUAAUAUGUUGCAAUGCAUGUUUCGGAUAACAAUGAGGCAAAAUUGAGUUAUUUAUUAACUCAA------------- ..((((((((...((((((..((((......))))..))))))(.(((((((......))))))).).))))))))..........((((((((.....))))))))------------- ( -29.70) >consensus AUUUGUUGUCGCUGAAUACAAUAUGGUGCAACAGGGCGUAUGAGUAAUAUGUUGCACAGUAUGUUGUGGAUAACAAUAAGCCAAAAUUGAGUUAUUCAUUAACUCAAUA_CAU_UAUGCU .((((((((..(((...(((.(((.((((((((..((......))....)))))))).))))))..)))...)))))))).....(((((((((.....)))))))))............ (-29.46 = -29.13 + -0.32)


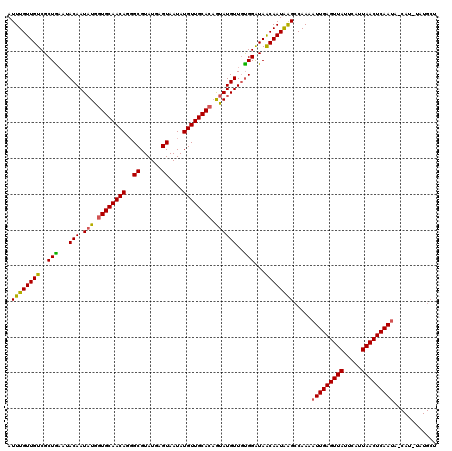
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:08:32 2006