
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,973,828 – 9,973,926 |
| Length | 98 |
| Max. P | 0.645145 |

| Location | 9,973,828 – 9,973,926 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 88.31 |
| Mean single sequence MFE | -30.33 |
| Consensus MFE | -23.14 |
| Energy contribution | -24.45 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.645145 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9973828 98 + 22224390 GUAUUAGUGUGCUAGAGUUGCAGUUGCACUUUGCGAUUCGUGA-AUGCAUUUUGGCCAUUGAUUGAAAUUGACCAAUUCGCUGUGGUGGCGCUAAUGCA------ ((((((((((((..(((((((((.......)))))))))....-..))......(((((.(((((........)))))....))))).)))))))))).------ ( -31.20) >DroSec_CAF1 19217 95 + 1 GUAUUAGUGUGCGAGAGUUGCAGUUGCACU-UGCGACUCGUGAAAUGCAUUUUGGCCAUUGAUUGAAAUUGACCAAUUCGCUGUGGUGGCGCA---GCA------ ..(((((((.((..(((((((((......)-))))))))((.....))......)))))))))................((((((....))))---)).------ ( -28.50) >DroEre_CAF1 6545 101 + 1 GUAUUAGUGUGC--GAACUGGAGUUGCACU-UGCGAUUCGUGA-AUGCAUUUUGGCCAUUGAUGGAAAUUGACCAAUUCGCUGUGGUGGCGCUAAUGCAACAACG ((((((((((((--(((.(((..((((...-.))))(((((.(-((((......).)))).)))))......))).))))).......))))))))))....... ( -30.61) >DroYak_CAF1 6782 101 + 1 GUAUUAGUGUGC--GACUUGCAGUGGCACU-UGCGAUUCGUGA-AUGCAUUUUGGCCAUUGAUUGAAAUUGACCAAUUCGCUGUGGUGGCGCUAAUGCAACAACA ((((((((..((--.(((.(((((((((.(-..((...))..)-.)))...(((((.(((......))).).))))..))))))))).))))))))))....... ( -31.00) >consensus GUAUUAGUGUGC__GAGUUGCAGUUGCACU_UGCGAUUCGUGA_AUGCAUUUUGGCCAUUGAUUGAAAUUGACCAAUUCGCUGUGGUGGCGCUAAUGCA______ ((((((((((((......((.((((((.....))))))))......))......(((((.(((((........)))))....))))).))))))))))....... (-23.14 = -24.45 + 1.31)


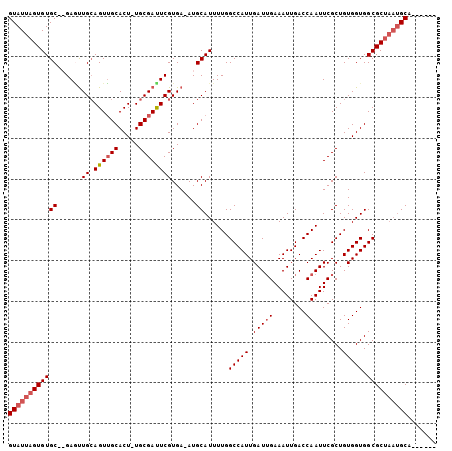
| Location | 9,973,828 – 9,973,926 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 88.31 |
| Mean single sequence MFE | -21.95 |
| Consensus MFE | -16.59 |
| Energy contribution | -17.34 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.614248 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9973828 98 - 22224390 ------UGCAUUAGCGCCACCACAGCGAAUUGGUCAAUUUCAAUCAAUGGCCAAAAUGCAU-UCACGAAUCGCAAAGUGCAACUGCAACUCUAGCACACUAAUAC ------((((((..(((.......)))..(((((((...........))))))))))))).-.........((..(((((....)).)))...)).......... ( -19.10) >DroSec_CAF1 19217 95 - 1 ------UGC---UGCGCCACCACAGCGAAUUGGUCAAUUUCAAUCAAUGGCCAAAAUGCAUUUCACGAGUCGCA-AGUGCAACUGCAACUCUCGCACACUAAUAC ------((.---((((........(((..(((((((...........)))))))..))).......((((.(((-........))).)))).))))))....... ( -21.80) >DroEre_CAF1 6545 101 - 1 CGUUGUUGCAUUAGCGCCACCACAGCGAAUUGGUCAAUUUCCAUCAAUGGCCAAAAUGCAU-UCACGAAUCGCA-AGUGCAACUCCAGUUC--GCACACUAAUAC (((...((((((..(((.......)))..(((((((...........))))))))))))).-..))).......-.(((((((....))).--))))........ ( -24.50) >DroYak_CAF1 6782 101 - 1 UGUUGUUGCAUUAGCGCCACCACAGCGAAUUGGUCAAUUUCAAUCAAUGGCCAAAAUGCAU-UCACGAAUCGCA-AGUGCCACUGCAAGUC--GCACACUAAUAC .((...((((((..(((.......)))..(((((((...........))))))))))))).-..))........-.((((.((.....)).--))))........ ( -22.40) >consensus ______UGCAUUAGCGCCACCACAGCGAAUUGGUCAAUUUCAAUCAAUGGCCAAAAUGCAU_UCACGAAUCGCA_AGUGCAACUGCAACUC__GCACACUAAUAC ......((((((..(((.......)))..(((((((...........)))))))))))))................((((.............))))........ (-16.59 = -17.34 + 0.75)

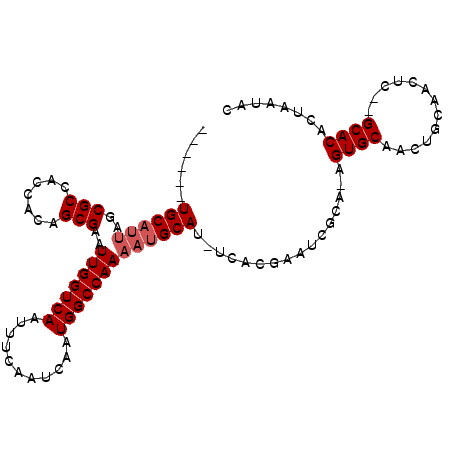
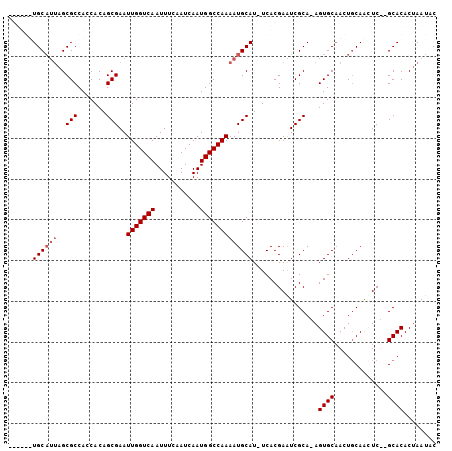
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:07:48 2006