
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,961,335 – 9,961,494 |
| Length | 159 |
| Max. P | 0.711662 |

| Location | 9,961,335 – 9,961,435 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 79.82 |
| Mean single sequence MFE | -13.98 |
| Consensus MFE | -11.94 |
| Energy contribution | -11.06 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.711662 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9961335 100 - 22224390 UUGCAGCAAA-AGUACAAGUAUCUUACAGAUACG----GCACAGCUUCUUGGCUAGAUAAACUUCAAAUAUAA-------ACAAAAAAAAA-AAAGAAGUAAAGAGGAAAAGC .(((.((...-.))....(((((.....))))).----))).((((....))))......(((((........-------...........-...)))))............. ( -12.25) >DroEre_CAF1 23923 113 - 1 GUGCAGCAAAAAGUACAAGUAUCUUACAGAUACGAACGAAACAGCUUCUUGGCUAGGUAAAAUCCAAAUAUAAGAGAGAGAUAAAAAAAAAAACAGAAGUGGAGAGGAAAAGU ((((........))))..(((((.....)))))...........((((((.(((.((......))...(((.........)))..............))).))))))...... ( -17.50) >DroYak_CAF1 23844 103 - 1 GUGCAGCAAA-AUUACAAGUAUCUUAUAGAUACG----AAACAGCUUCUUCACUAGGUAAAAUCCAAAUA----AGAAAGAUAGAAAAAAA-AAAGAGGUAGAGAGGAAAAGC .....((...-.......(((((.....))))).----......(((((..(((.((......))..((.----......)).........-.....)))..)))))....)) ( -12.20) >consensus GUGCAGCAAA_AGUACAAGUAUCUUACAGAUACG____AAACAGCUUCUUGGCUAGGUAAAAUCCAAAUAUAA_AGA_AGAUAAAAAAAAA_AAAGAAGUAGAGAGGAAAAGC .....((...........(((((.....)))))...........((((((.(((...........................................))).))))))....)) (-11.94 = -11.06 + -0.88)



| Location | 9,961,400 – 9,961,494 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 75.52 |
| Mean single sequence MFE | -14.70 |
| Consensus MFE | -12.68 |
| Energy contribution | -11.80 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.21 |
| Mean z-score | -0.59 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.590789 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9961400 94 + 22224390 GC----CGUAUCUGUAAGAUACUUGUACU-UUUGCUGCAAUGACAUUUAAUUGU-UGCAAUACACCUGCGGAAAAAAGAUGUAUAUUUAAAACCUGAACA .(----((((((.....))))).((((..-.((((.(((((........)))))-.)))))))).....))............................. ( -15.90) >DroEre_CAF1 23996 88 + 1 UUCGUUCGUAUCUGUAAGAUACUUGUACUUUUUGCUGCACUGGCAUUUAAUUGU-UGCAACACACCUGCAAAAUUUGA-----------AAAACUAAAUA .......(((((.....)))))..........((((.....))))(((((...(-((((.......)))))...))))-----------).......... ( -13.80) >DroYak_CAF1 23912 87 + 1 UU----CGUAUCUAUAAGAUACUUGUAAU-UUUGCUGCACUGACAUUUAAUUGUUUGCAACACACCUGCAAAAAAAAGAUG--------UAAACUGAAUA ((----((((((.....)))))(((((..-..((.((((..((((......)))))))).))....)))))..........--------......))).. ( -14.40) >consensus UU____CGUAUCUGUAAGAUACUUGUACU_UUUGCUGCACUGACAUUUAAUUGU_UGCAACACACCUGCAAAAAAAAGAUG________AAAACUGAAUA .......(((((.....)))))(((((.....((.((((...(((......))).)))).))....)))))............................. (-12.68 = -11.80 + -0.88)

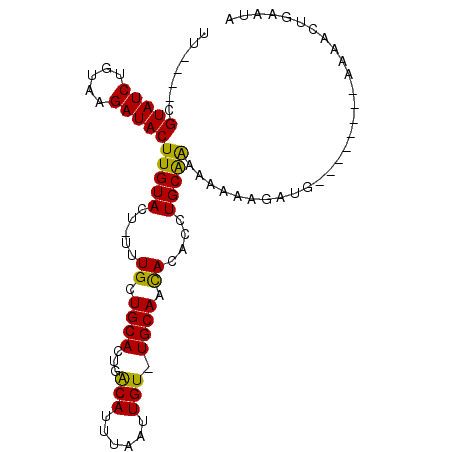

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:07:36 2006