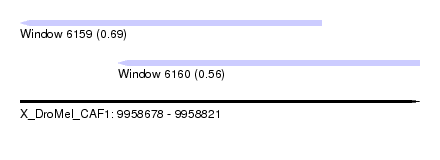
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,958,678 – 9,958,821 |
| Length | 143 |
| Max. P | 0.685721 |

| Location | 9,958,678 – 9,958,786 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 67.35 |
| Mean single sequence MFE | -37.35 |
| Consensus MFE | -12.06 |
| Energy contribution | -13.25 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.32 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685721 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9958678 108 - 22224390 CCCAGAGCGG-------CCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCUUGGUUAUAUAGCCGCCGAUACACACACACAUAUUUACA-GAACGUA----UGUGUAUACGUAUACAGCUC ....((((((-------(((.(.(((....))).).)))))....((((..((((....))))))))((((....((((((((.....-....)))----)))))....))))...)))) ( -36.10) >DroSec_CAF1 26656 104 - 1 CCCAGAGCGG-------CCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCAUGGCUAUAUAGCCGCCGAUACAC----ACGUAUUUACA-GUAUGUA----UGUAGANNNNNNNNNNNNNN ....((..((-------(((.(.(((....))).).))))).)).((((..((((....)))))))).((((.----((((((.....-)))))).----))))................ ( -34.10) >DroEre_CAF1 21286 106 - 1 CCCAGAGCGGCCAGCGGCCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCCUGGCUAUUU--------AUAUUUA--UAUGUUUUUACUUGUAUAUA----UGUAUGUGUGCGUACAGCUC ....(((((((((..((((.(((((((.........)))))))...)))))))))....--------((((.((--((.((....)).)))).)))----)(((((....))))).)))) ( -33.90) >DroYak_CAF1 21062 104 - 1 CCCAGAGCGG--------------CCAAGUGGCAAGUGGCUAUCAUGGCCUGGCUAUAUGCCCGCCGAUACACA--CGUAUUUGUACUUGUGUAUAAGUAUGUAUGUAUGUAUGCCGCUC ....((((((--------------(....((((....((((.....)))).(((.....))).))))(((((.(--(((((((((((....)))))))))))).)))))....))))))) ( -45.30) >consensus CCCAGAGCGG_______CCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCCUGGCUAUAUAGCCGCCGAUACACA__CACAUAUUUACA_GUAUAUA____UGUAUAUAUGUAUACAGCUC ....((((............(((((((...(((..(((.....))).))))))))))..........(((((.....((((..........)))).....)))))...........)))) (-12.06 = -13.25 + 1.19)

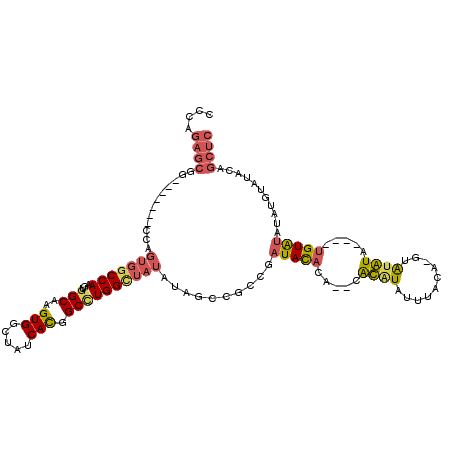
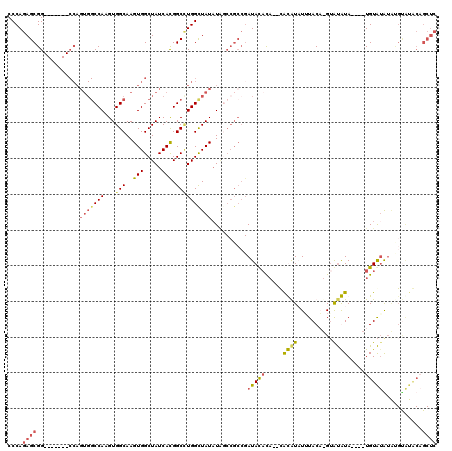
| Location | 9,958,713 – 9,958,821 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.34 |
| Mean single sequence MFE | -38.05 |
| Consensus MFE | -24.92 |
| Energy contribution | -26.05 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.561227 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9958713 108 - 22224390 GAAUUUGGCCAGGUAAGCCAGGUAGG-----UAAGUGCAGCCCAGAGCGG-------CCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCUUGGUUAUAUAGCCGCCGAUACACACACACA .....(((((..(((.(((.....))-----)...))).((.....))))-------)))((((((....)))..(((..((((.(((((.........)))))..))))..))).))). ( -36.70) >DroSec_CAF1 26691 104 - 1 GAAUUUGGCCAGGUAAGCCAGGUAGG-----UAAGUGCAGCCCAGAGCGG-------CCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCAUGGCUAUAUAGCCGCCGAUACAC----ACG ..(((((((.......)))))))...-----...(((.......((..((-------(((.(.(((....))).).))))).)).((((..((((....))))))))...)))----... ( -36.30) >DroEre_CAF1 21322 110 - 1 GAAUUUGGCCAAGAAAGCCAGGUAAGCCACGUAAGUGCAGCCCAGAGCGGCCAGCGGCCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCCUGGCUAUUU--------AUAUUUA--UAUG ...........(((.((((((((..(((((......((.(((......)))..))((((...))))..)))))..(((.....))).)))))))).)))--------.......--.... ( -39.00) >DroYak_CAF1 21102 100 - 1 GAAUUUGGCCAGGUAAGCCAGGCAAGCCA----AGUGCAGCCCAGAGCGG--------------CCAAGUGGCAAGUGGCUAUCAUGGCCUGGCUAUAUGCCCGCCGAUACACA--CGUA ....(((((..((((((((((((.(((((----..(((.(((......))--------------)......)))..)))))......))))))))...)))).)))))......--.... ( -40.20) >consensus GAAUUUGGCCAGGUAAGCCAGGUAAG_____UAAGUGCAGCCCAGAGCGG_______CCAGUGGCCAAGUGGCAAGUGGCUAUCACGGCCUGGCUAUAUAGCCGCCGAUACACA__CACA ....(((((...(((((((((((...........(((.(((((.....(((.....)))....(((....)))..).))))..))).)))))))).)))....)))))............ (-24.92 = -26.05 + 1.13)

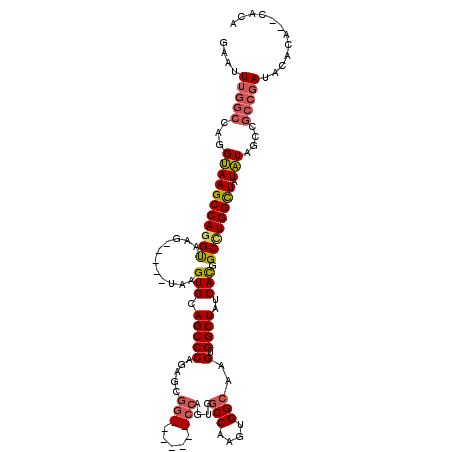

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:07:30 2006