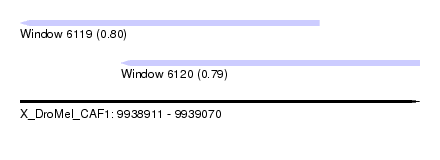
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,938,911 – 9,939,070 |
| Length | 159 |
| Max. P | 0.801075 |

| Location | 9,938,911 – 9,939,030 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.28 |
| Mean single sequence MFE | -42.00 |
| Consensus MFE | -35.12 |
| Energy contribution | -35.00 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801075 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
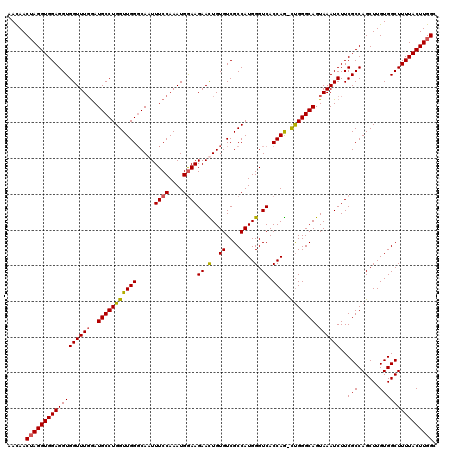
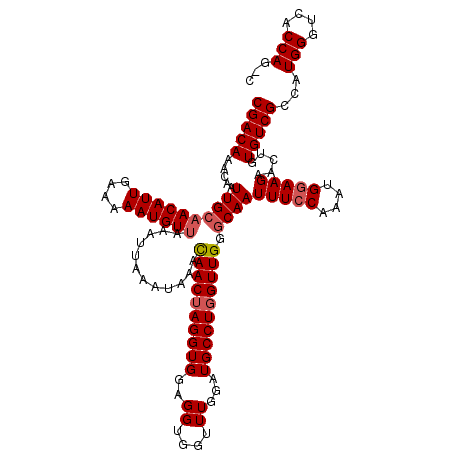
>X_DroMel_CAF1 9938911 119 - 22224390 AACAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGGCAAUUUCCAAAUGAAAGAACUGUGUCGCCAUGGGUCACCAG-UUGGGCAGUAAAUCUUCGCCAGCUUGUGGCUUUUACUUGGC .....(((((((((((((((((((((((((....)))))...))))))).............))))...((((((.((-(((((.((.....)).).)))))).))))))))))))))). ( -40.11) >DroSec_CAF1 8686 120 - 1 AACAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGGCAAUUUCCAAAUGGAAGAAUUGUGUCGCCAUGGGUCACCAACUUGGGCAGUAAAUCUUCGCCAGCUUGUGGCUUUUACUUGGC .....((((((((((((((((((..((((((((.(.(((((((((....)))..)))))).).))).(((....)))....))))).))))))..(((......))))))))))))))). ( -39.50) >DroEre_CAF1 8385 119 - 1 AAUAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGCCAAUUUCCAAAUGGAAGAACUGUGUCGCCAUGGGUCACCAG-CUGGGCAGUAAAUCUUCGCCAGCUUGUGGCUUUUACUUGGG .....((((((((((((((((((..((((..(((((.....((((....))))((.(((((....))))).)).))))-)..)))).))))))..(((......))))))))))))))). ( -44.10) >DroYak_CAF1 8797 119 - 1 AAUAACAAGGUGGAGGUGGUUUCAAUGCCUGGUUGGGCAAUUUCCAAAUGGAAGAAUUGUGUCGCCAUGGGUCACCAG-CCGGGCAGUAAAUCUUCGCCAGCUUGUGGCUUUUACUUGGA ....(((((.(((.((.(((((...((((((((((((((((((((....)))..))))))...(((...)))..))))-)))))))..))))).)).))).))))).............. ( -44.30) >consensus AACAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGGCAAUUUCCAAAUGGAAGAACUGUGUCGCCAUGGGUCACCAG_CUGGGCAGUAAAUCUUCGCCAGCUUGUGGCUUUUACUUGGC .....((((((((((((((((((..(((((((((((.....((((....))))((.(..((....))..).)).)))).))))))).))))))..(((......))))))))))))))). (-35.12 = -35.00 + -0.12)


| Location | 9,938,951 – 9,939,070 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.98 |
| Mean single sequence MFE | -31.10 |
| Consensus MFE | -27.05 |
| Energy contribution | -27.80 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792475 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
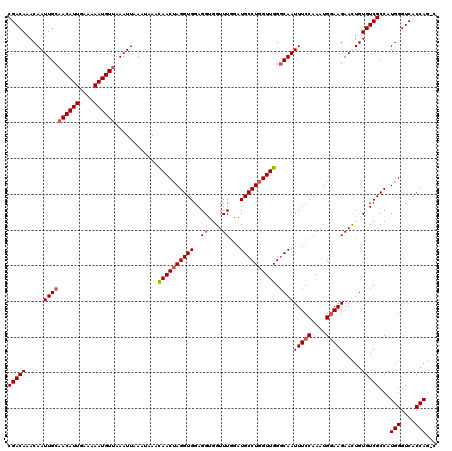
>X_DroMel_CAF1 9938951 119 - 22224390 CGACAAACAAUUGCAACAUUGAAAAAUGUUAAAUUAAAUAAACAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGGCAAUUUCCAAAUGAAAGAACUGUGUCGCCAUGGGUCACCAG-U .(((....((((..((((((....)))))).))))..........((((((((.(((.((((((((((((....)))))...)))))))......)))...))))).)))))).....-. ( -31.60) >DroSec_CAF1 8726 120 - 1 CGACAAACAAUUGCAACAUUGAAAAAUGUUAAAUUAAAUAAACAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGGCAAUUUCCAAAUGGAAGAAUUGUGUCGCCAUGGGUCACCAACU ((((..((((((((((((((....))))))............((((((((((..((....))...)))))))))).))..(((((....)))))))))))))))...(((....)))... ( -33.50) >DroEre_CAF1 8425 119 - 1 CGACAAACAAUUGCAACAUUGAAAAAUGUAAAAUUAAAUAAAUAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGCCAAUUUCCAAAUGGAAGAACUGUGUCGCCAUGGGUCACCAG-C .(((....((((...(((((....)))))..))))..........((((((((..(..((((....(((.....)))....((((....))))))))..).))))).)))))).....-. ( -28.30) >DroYak_CAF1 8837 119 - 1 CGACAAACAAUUGCAACAUUGAAAAAUGUUAAAUUAAAUAAAUAACAAGGUGGAGGUGGUUUCAAUGCCUGGUUGGGCAAUUUCCAAAUGGAAGAAUUGUGUCGCCAUGGGUCACCAG-C ........((((..((((((....)))))).)))).............(((((..(((((..((.(((((....))))).(((((....))))).....))..)))))...)))))..-. ( -31.00) >consensus CGACAAACAAUUGCAACAUUGAAAAAUGUUAAAUUAAAUAAACAACUAGGUGGAGGUGGUUUGGAUGCCUGGUUGGGCAAUUUCCAAAUGGAAGAACUGUGUCGCCAUGGGUCACCAG_C (((((.....((((((((((....))))))............((((((((((..((....))...)))))))))).))))(((((....))))).....)))))...(((....)))... (-27.05 = -27.80 + 0.75)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:53 2006