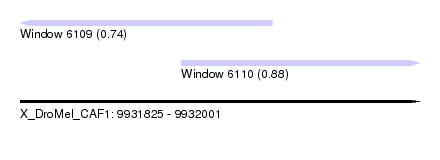
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,931,825 – 9,932,001 |
| Length | 176 |
| Max. P | 0.875136 |

| Location | 9,931,825 – 9,931,936 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.47 |
| Mean single sequence MFE | -22.73 |
| Consensus MFE | -16.11 |
| Energy contribution | -15.54 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744407 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
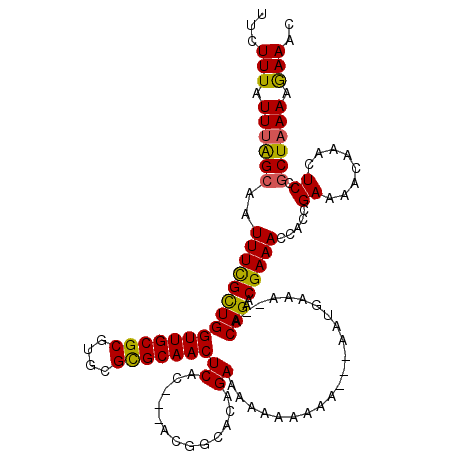
>X_DroMel_CAF1 9931825 111 - 22224390 UUCUUUAUUUAGCAAUUUUGUUGGUUGCGCGUGCGCGCAACUCAC---ACGGCACAGAAAA----AAAAGAAUGAAA--ACAGAACGAAACCAACGAAAACAAACUCCGCUAAAAGAAAC ((((((...((((.....(((.((((((((....)))))))).))---)............----............--......((.......))............)))))))))).. ( -22.40) >DroSec_CAF1 1424 115 - 1 UUCUUUAUUUAGCAAUUUCGCUGGUUGCGCGUGCGCGCAACUCAC---ACGGCACAGAAAAAAAAAAAAGAAUGAAA--ACAGAACGAAACCACCGAAAACAAACUCCGCUAAAAGAAAC ((((((...((((......(((((((((((....)))))))....---.))))........................--......((.......))............)))))))))).. ( -22.80) >DroEre_CAF1 1445 117 - 1 UUCUUUAUUUAGAAAUUUCGCUGGUUGCGCGUGCGUGCAACUCACUGCAAUGCACAGAAAAAAAAAA---GAGGGCACCACAGAACGAAACCACCGAAAAAAAACUCCGCUUAAAAAAAC ((((......)))).(((((.(((((.((.(((((((((......))).))))))............---..((...))......)).))))).)))))..................... ( -24.20) >DroYak_CAF1 1508 112 - 1 UUCUUUUUUUGGCAAUUUUGCUGGUUGCGCGUGCGUGCAACUCAC---ACUGCACAGAAAAAAAAAA---AACGAAA--ACAGAACGAAACCACCGACAACAAACUCCGCUAAAAGAAAC ...((((((((((.....((((((((((((....))))))).)).---...))).............---.......--......((.......))............)))))))))).. ( -21.50) >consensus UUCUUUAUUUAGCAAUUUCGCUGGUUGCGCGUGCGCGCAACUCAC___ACGGCACAGAAAAAAAAAA___AAUGAAA__ACAGAACGAAACCACCGAAAACAAACUCCGCUAAAAGAAAC ...(((.((((((..(((((((((((((((....)))))))((.............))......................)))..))))).....((........)).)))))).))).. (-16.11 = -15.54 + -0.56)

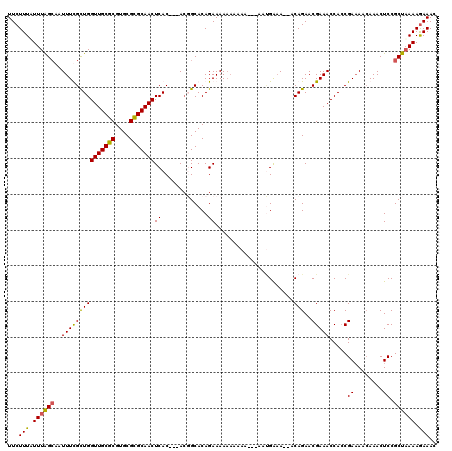
| Location | 9,931,896 – 9,932,001 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 87.61 |
| Mean single sequence MFE | -35.83 |
| Consensus MFE | -31.63 |
| Energy contribution | -32.30 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.875136 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
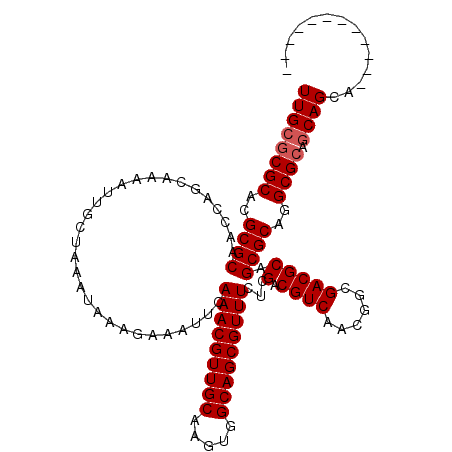
>X_DroMel_CAF1 9931896 105 + 22224390 UUGCGCGCACGCGCAACCAACAAAAUUGCUAAAUAAAGAAAUUCAAACGUUGCAAGUGGCAGCGUUUCUCGACGUCAACGGCGACGCAGCGCAGGCGCAGCAGCA------------ ((((((((..((((..............................(((((((((.....)))))))))...(.((((......))))).))))..)))).))))..------------ ( -35.20) >DroSec_CAF1 1499 105 + 1 UUGCGCGCACGCGCAACCAGCGAAAUUGCUAAAUAAAGAAAUUCAAACGUUGCAAGUGGCAGCGUUUCUCGACGUCAACGGCGACGCAGCGCAGGCGCAGCAGCA------------ ((((((((..((((....((((....)))).................((((((...((((..((.....))..))))...))))))..))))..)))).))))..------------ ( -36.90) >DroYak_CAF1 1580 117 + 1 UUGCACGCACGCGCAACCAGCAAAAUUGCCAAAAAAAGAAAUACAAACGUUGCAAGUGGCAGCGUUUUUCGACGUCGACGGCGACGCAGCGCAGGCGCAACAGCAGCAGUAGCAGCA ((((((((..((((.....(((....)))........((((....((((((((.....))))))))))))(.(((((....)))))).))))..))((....))....)).)))).. ( -35.40) >consensus UUGCGCGCACGCGCAACCAGCAAAAUUGCUAAAUAAAGAAAUUCAAACGUUGCAAGUGGCAGCGUUUCUCGACGUCAACGGCGACGCAGCGCAGGCGCAGCAGCA____________ ((((((((..((((..............................(((((((((.....)))))))))...(.((((......))))).))))..)))).)))).............. (-31.63 = -32.30 + 0.67)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:44 2006