
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,923,516 – 9,923,608 |
| Length | 92 |
| Max. P | 0.676483 |

| Location | 9,923,516 – 9,923,608 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 82.21 |
| Mean single sequence MFE | -23.60 |
| Consensus MFE | -10.57 |
| Energy contribution | -10.69 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.676483 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
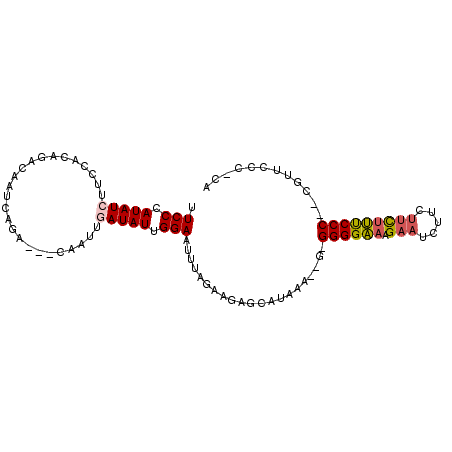
>X_DroMel_CAF1 9923516 92 + 22224390 UUCCCAUAUCUUCCUCAGACAAUCAGA---CAAUUGAUAUUGGAAUUUGGCAGAGCAUAGUGGGAGGGGAAAGAAUCAU---CUUUCCC--CGUUCCA-CA ((((.(((((....((.(.....).))---.....))))).))))..............(((((((((((((((....)---)))))))--).)))))-). ( -30.30) >DroSec_CAF1 20709 93 + 1 UUCCCAUAUCUUCCACAGACAAUCAGA---CAGUUAAUAUUGGAAUUUAGGAGAGCAUAAA--GUGGGGAAAGAAUCUUCUUCUUUCCC--CGUUCCC-CA ......((..(((((..(((.......---..))).....)))))..))((.((((.....--..((((((((((.....)))))))))--))))).)-). ( -22.71) >DroSim_CAF1 20891 93 + 1 UUCCCAUAUCUUCCACAGACAAUCAGA---CAGUUAAUAUUGGAAUUCAGGAGAGCAUAGA--GUGGGGAAAGAAUCUUCUUCUUUCCC--CGUUCCC-CA ((((......(((((..(((.......---..))).....)))))....))))......((--((((((((((((.....)))))))))--)))))..-.. ( -22.90) >DroEre_CAF1 21141 92 + 1 CUCCCAUAUCUUCCUCAGACAAUCAGA---CAAUUGAUAUUGGAUUUUAGAAGAGCAUAAU--G-GGGGCAAGAAUCUACUUCUGCCCC--CUUUCCC-CA .(((.(((((....((.(.....).))---.....))))).))).................--(-((((((.(((.....)))))))))--)......-.. ( -21.80) >DroYak_CAF1 20904 98 + 1 CUCCCAUAUCUUCCGCAGACAAUCAGACGACAAUUGAUAUUGGAUUUUAGAAAAGCACAAA--G-GGGGCAAGAAUCUCCUUUCGCCCCCCCUUUCCCACC .(((.(((((.........................))))).)))..............(((--(-((((...(((......)))...))))))))...... ( -20.31) >consensus UUCCCAUAUCUUCCACAGACAAUCAGA___CAAUUGAUAUUGGAAUUUAGAAGAGCAUAAA__G_GGGGAAAGAAUCUUCUUCUUUCCC__CGUUCCC_CA .(((.(((((.........................))))).))).....................((((((.(((.....)))))))))............ (-10.57 = -10.69 + 0.12)


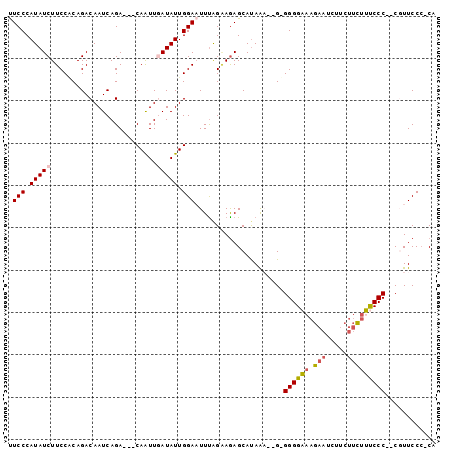
| Location | 9,923,516 – 9,923,608 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 82.21 |
| Mean single sequence MFE | -28.22 |
| Consensus MFE | -14.22 |
| Energy contribution | -15.74 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.536341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9923516 92 - 22224390 UG-UGGAACG--GGGAAAG---AUGAUUCUUUCCCCUCCCACUAUGCUCUGCCAAAUUCCAAUAUCAAUUG---UCUGAUUGUCUGAGGAAGAUAUGGGAA .(-(((...(--(((((((---(....)))))))))..)))).........(((..((((.....(((((.---...))))).....))))....)))... ( -29.50) >DroSec_CAF1 20709 93 - 1 UG-GGGAACG--GGGAAAGAAGAAGAUUCUUUCCCCAC--UUUAUGCUCUCCUAAAUUCCAAUAUUAACUG---UCUGAUUGUCUGUGGAAGAUAUGGGAA .(-((((..(--(((((((((.....)))))))))).(--.....).)))))....(((((..((((....---..))))(((((.....)))))))))). ( -26.70) >DroSim_CAF1 20891 93 - 1 UG-GGGAACG--GGGAAAGAAGAAGAUUCUUUCCCCAC--UCUAUGCUCUCCUGAAUUCCAAUAUUAACUG---UCUGAUUGUCUGUGGAAGAUAUGGGAA ((-(((...(--(((((((((.....)))))))))).)--))))...((((.((..(((((....((((..---...).)))....)))))..)).)))). ( -27.40) >DroEre_CAF1 21141 92 - 1 UG-GGGAAAG--GGGGCAGAAGUAGAUUCUUGCCCC-C--AUUAUGCUCUUCUAAAAUCCAAUAUCAAUUG---UCUGAUUGUCUGAGGAAGAUAUGGGAG .(-(((...(--(((((((((.....))).))))))-)--......)))).......(((.(((((..((.---((.(.....).)).)).))))).))). ( -26.60) >DroYak_CAF1 20904 98 - 1 GGUGGGAAAGGGGGGGCGAAAGGAGAUUCUUGCCCC-C--UUUGUGCUUUUCUAAAAUCCAAUAUCAAUUGUCGUCUGAUUGUCUGCGGAAGAUAUGGGAG ..(((((((((((((((((.((....)).)))))))-)--).....))))))))...(((.(((((..((((.(.(.....).).))))..))))).))). ( -30.90) >consensus UG_GGGAACG__GGGAAAGAAGAAGAUUCUUUCCCC_C__UUUAUGCUCUCCUAAAUUCCAAUAUCAAUUG___UCUGAUUGUCUGAGGAAGAUAUGGGAA ............(((((((((.....)))))))))..............(((((..((((.....(((((.......))))).....))))....))))). (-14.22 = -15.74 + 1.52)

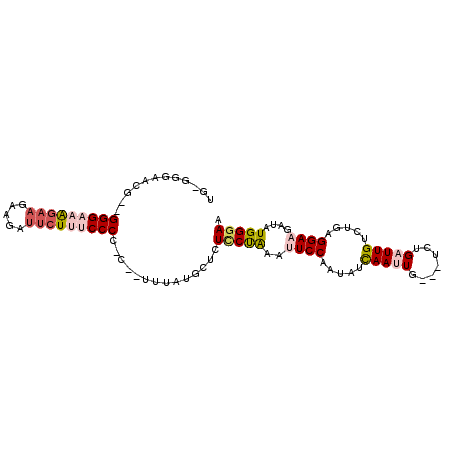

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:38 2006