
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,914,217 – 9,914,335 |
| Length | 118 |
| Max. P | 0.993830 |

| Location | 9,914,217 – 9,914,311 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 91.27 |
| Mean single sequence MFE | -22.22 |
| Consensus MFE | -19.94 |
| Energy contribution | -20.00 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.35 |
| SVM RNA-class probability | 0.992826 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
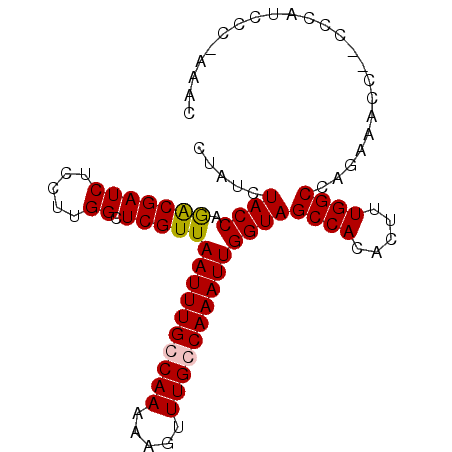
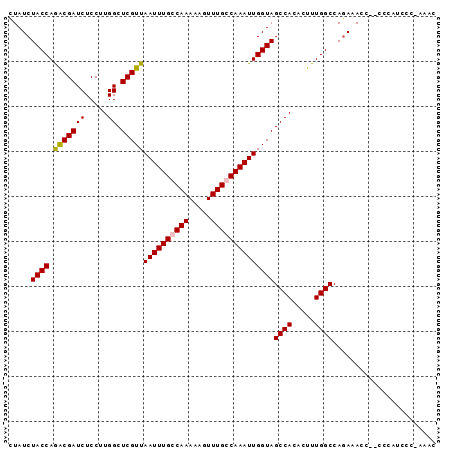
>X_DroMel_CAF1 9914217 94 + 22224390 UUAUCUACCAGGCGAUCUCCUUGGCUCGUUAAUUUGCCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACCUCCCCACCCC-AAGC .....((((((((((.((..(((((..........)))))..)).)))))....)))))((((.....))))..................-.... ( -23.10) >DroSec_CAF1 11663 92 + 1 CUAUCUACCAAACGAUCUCCUUGGCUCGUUAAUUUGCCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACC--CCCAUCCC-AAAC .....((((((.(((.((..(((((..........)))))..)).))).....))))))((((.....))))........--........-.... ( -16.70) >DroSim_CAF1 11840 92 + 1 CUAUCUACCAGGCGAUCUCCUUGGCUCGUUAAUUUGGCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACC--CCCAUCCC-AAAC ..........(((.(......(((((.(..((((((((((.....))))))))))..)))))).....).))).......--........-.... ( -25.40) >DroYak_CAF1 11632 93 + 1 CUAUCUACCAGACGAUCUCCUUGGCUCGUUAAUUUGGCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAAACACA--CCAAACCGAAAAU .....((((.(((((((.....)).)))))((((((((((.....))))))))))))))((((.....))))........--............. ( -23.70) >consensus CUAUCUACCAGACGAUCUCCUUGGCUCGUUAAUUUGCCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACC__CCCAUCCC_AAAC .....((((.(((((((.....)).)))))((((((((((.....))))))))))))))((((.....))))....................... (-19.94 = -20.00 + 0.06)

| Location | 9,914,244 – 9,914,335 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 91.71 |
| Mean single sequence MFE | -18.78 |
| Consensus MFE | -17.61 |
| Energy contribution | -17.93 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.94 |
| SVM decision value | 2.43 |
| SVM RNA-class probability | 0.993830 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
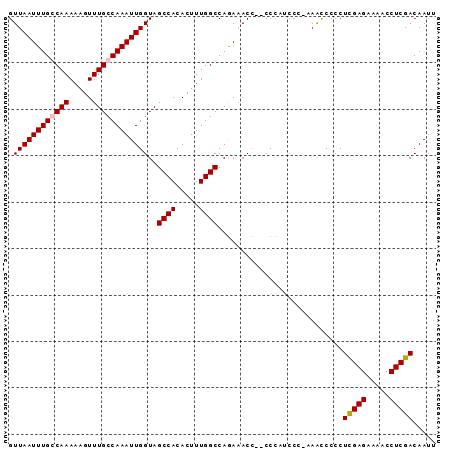
>X_DroMel_CAF1 9914244 91 + 22224390 GUUAAUUUGCCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACCUCCCCACCCC-AAGCUCCCUCGAGAAAACCUCGACAAUU ........((((((..(((((((.....)))))..))..))))))..................-........(((((.....)))))..... ( -16.80) >DroSec_CAF1 11690 89 + 1 GUUAAUUUGCCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACC--CCCAUCCC-AAACCCCCUCGAGAAAACCUCGACAAUU ........((((((..(((((((.....)))))..))..))))))........--........-........(((((.....)))))..... ( -16.80) >DroSim_CAF1 11867 89 + 1 GUUAAUUUGGCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACC--CCCAUCCC-AAACCCCCUCGAGAAAACCUCGACAAUU .((((((((((((.....))))))))))))..((((.....))))........--........-........(((((.....)))))..... ( -21.80) >DroYak_CAF1 11659 90 + 1 GUUAAUUUGGCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAAACACA--CCAAACCGAAAAUCCCCUUGAGAAAACCUCGACAAUU .((((((((((((.....))))))))))))..((((.....))))........--.................(((((.....)))))..... ( -19.70) >consensus GUUAAUUUGCCAAAAAGUUUGCCAAAUUGGUAGCCACACUUUGGCCAGAAACC__CCCAUCCC_AAACCCCCUCGAGAAAACCUCGACAAUU .((((((((((((.....))))))))))))..((((.....))))...........................(((((.....)))))..... (-17.61 = -17.93 + 0.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:32 2006