
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,900,484 – 9,900,585 |
| Length | 101 |
| Max. P | 0.947343 |

| Location | 9,900,484 – 9,900,585 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -27.40 |
| Consensus MFE | -23.60 |
| Energy contribution | -23.36 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.947343 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
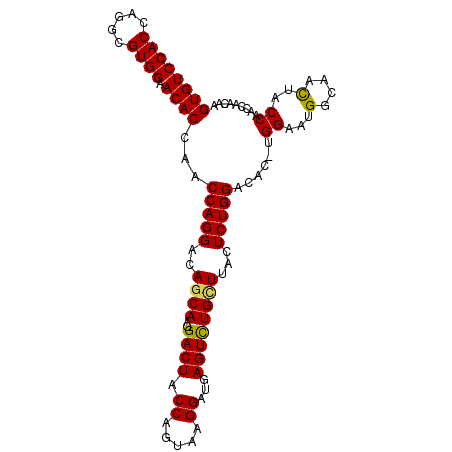
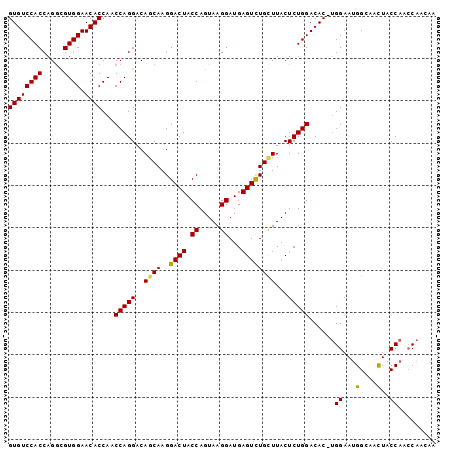
>X_DroMel_CAF1 9900484 101 + 22224390 GUGUCCACCAGGCGUGGAACACCAACCAGGACAGCAAGGACUACCAGUAAGGAUGAGUCUGCUUACUCUGGACACCUGGAAUGGCAACUACCAAACAACAA .((((..(((((.((((....))..(((((..((((..((((.((.....))...))))))))...))))))).)))))...))))............... ( -29.00) >DroSec_CAF1 4776 100 + 1 GUGUCCACCAGGUGUGGAACACCAACCAGGACAGCAAGGACUACCAGUAAGGAUGAGUCUGCUUACUCUGGACAC-UGGAAAGGCAACUACCAACCAAAAA (((((((((.(((.(((....)))))).))..((((..((((.((.....))...)))))))).....)))))))-(((..((....)).)))........ ( -30.10) >DroSim_CAF1 4800 100 + 1 GUGUCCACCAGGCGUGGAACACCAACCAGGACAGCAAGGACUACCAGUAAGGAUGAGUCUGUUUACUCUGGACAC-UGGAAUGGCAACUACCAACCAAAAA (((((((((.((..(((....))).)).)).......(((((.((.....))...)))))........)))))))-(((..(((......))).))).... ( -26.50) >DroEre_CAF1 4718 100 + 1 GUGUCCACCAGACGUGGAACACCAACCAGGACAACAAGGACUACCAGUAAGGAUGAGUCUGCUUACUCUGGACAC-CGGAAUGGCAACUACCAACCAACAA ((((((((.....)))).))))......((.......(((((.((.....))...)))))(((...(((((...)-))))..))).....))......... ( -24.90) >DroYak_CAF1 4850 100 + 1 GUGUCCACCAGGCGUGGAACACCAACCAGGACAGCAAGGACUACCAGUAAGGAUGAGUUUGCUUACUCUGGACAC-UGGAAUGGCAAUUACCAACCAACAA (((((((((.((..(((....))).)).))..((((..((((.((.....))...)))))))).....)))))))-(((..(((......))).))).... ( -26.50) >consensus GUGUCCACCAGGCGUGGAACACCAACCAGGACAGCAAGGACUACCAGUAAGGAUGAGUCUGCUUACUCUGGACAC_UGGAAUGGCAACUACCAACCAACAA ((((((((.....)))).))))...(((((..((((..((((.((.....))...))))))))...)))))......((...(....)..))......... (-23.60 = -23.36 + -0.24)

| Location | 9,900,484 – 9,900,585 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -31.34 |
| Consensus MFE | -28.08 |
| Energy contribution | -28.04 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.690230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
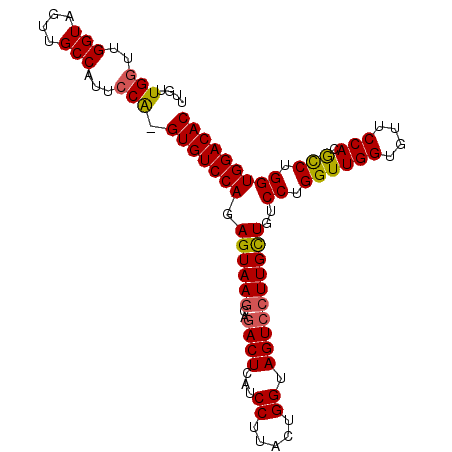
>X_DroMel_CAF1 9900484 101 - 22224390 UUGUUGUUUGGUAGUUGCCAUUCCAGGUGUCCAGAGUAAGCAGACUCAUCCUUACUGGUAGUCCUUGCUGUCCUGGUUGGUGUUCCACGCCUGGUGGACAC .((((...((((....))))..((((((((((((((((((..((((...((.....)).))))))))))...))))..((....))))))))))..)))). ( -32.70) >DroSec_CAF1 4776 100 - 1 UUUUUGGUUGGUAGUUGCCUUUCCA-GUGUCCAGAGUAAGCAGACUCAUCCUUACUGGUAGUCCUUGCUGUCCUGGUUGGUGUUCCACACCUGGUGGACAC ....(((..(((....)))...)))-(((((((.((((((..((((...((.....)).))))))))))..((.((((((....))).))).))))))))) ( -33.70) >DroSim_CAF1 4800 100 - 1 UUUUUGGUUGGUAGUUGCCAUUCCA-GUGUCCAGAGUAAACAGACUCAUCCUUACUGGUAGUCCUUGCUGUCCUGGUUGGUGUUCCACGCCUGGUGGACAC ....(((.((((....))))..)))-(((((((.(((((...((((...((.....)).)))).)))))..((.((((((....))).))).))))))))) ( -32.00) >DroEre_CAF1 4718 100 - 1 UUGUUGGUUGGUAGUUGCCAUUCCG-GUGUCCAGAGUAAGCAGACUCAUCCUUACUGGUAGUCCUUGUUGUCCUGGUUGGUGUUCCACGUCUGGUGGACAC ...((((.((((....))))..)))-)...((((..(((.((((((...((.....)).))))..)))))..))))...((((.((((.....)))))))) ( -26.60) >DroYak_CAF1 4850 100 - 1 UUGUUGGUUGGUAAUUGCCAUUCCA-GUGUCCAGAGUAAGCAAACUCAUCCUUACUGGUAGUCCUUGCUGUCCUGGUUGGUGUUCCACGCCUGGUGGACAC ....(((.((((....))))..)))-(((((((.((((((...(((...((.....)).))).))))))..((.((((((....))).))).))))))))) ( -31.70) >consensus UUGUUGGUUGGUAGUUGCCAUUCCA_GUGUCCAGAGUAAGCAGACUCAUCCUUACUGGUAGUCCUUGCUGUCCUGGUUGGUGUUCCACGCCUGGUGGACAC ....(((..(((....)))...))).(((((((.((((((..((((...((.....)).))))))))))..((.((((((....))).))).))))))))) (-28.08 = -28.04 + -0.04)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:24 2006